Engineering the Future of Cell Therapy: How Genetic Modification Supercharges MSC Immunomodulation
This article provides a comprehensive analysis for researchers and drug development professionals on leveraging genetic engineering to enhance the immunomodulatory potency of Mesenchymal Stromal Cells (MSCs).
Engineering the Future of Cell Therapy: How Genetic Modification Supercharges MSC Immunomodulation
Abstract
This article provides a comprehensive analysis for researchers and drug development professionals on leveraging genetic engineering to enhance the immunomodulatory potency of Mesenchymal Stromal Cells (MSCs). We first establish the foundational biology of native MSC immunomodulation and its limitations. We then detail current methodological approaches, including viral and non-viral gene delivery strategies targeting key pathways like IDO, TGF-β, and PD-L1. The discussion extends to critical troubleshooting of safety, persistence, and manufacturing challenges. Finally, we compare the efficacy of different genetic strategies against naive and pharmacologically enhanced MSCs, evaluating validation models from in vitro assays to clinical trials. This synthesis aims to guide the rational design of next-generation, engineered MSC therapies for autoimmune and inflammatory diseases.
The Blueprint of Immune Control: Understanding Native MSC Mechanisms and Therapeutic Limits
Within the broader thesis on Genetic modification of MSCs to enhance immunomodulation, it is foundational to first delineate the innate mechanisms of naive, unmodified Mesenchymal Stromal Cells (MSCs). Their potent immunomodulatory effects are not constitutive but are primarily licensed by inflammatory cues, mediated via two core modalities: paracrine signaling (soluble factors) and direct cell-cell contact. This application note provides a detailed experimental framework to dissect these mechanisms, serving as a critical baseline for subsequent studies on genetically engineered MSC variants (e.g., IDO1-overexpressing, PD-L1-modified cells).
Paracrine Signaling: Key Soluble Mediators and Protocols
Naive MSCs secrete a plethora of immunomodulatory molecules in response to inflammatory cytokines (primarily IFN-γ alone or in synergy with TNF-α, IL-1α, or IL-1β). The key mediators include Indoleamine 2,3-dioxygenase (IDO), Prostaglandin E2 (PGE2), Transforming Growth Factor-beta (TGF-β), and Interleukin-10 (IL-10).
Table 1: Core Paracrine Mediators from Inflammatory-Primed MSCs
| Mediator | Inducing Signal(s) | Primary Immune Target | Quantitative Range (in vitro) | Key Effect |
|---|---|---|---|---|
| IDO Activity | IFN-γ, TNF-α | T cells, NK cells | 50-95% Tryptophan depletion in 72h culture | Tryptophan catabolism, T cell cycle arrest/apoptosis, Treg induction |
| PGE2 | IFN-γ, TNF-α, IL-1β | Macrophages, T cells, DCs | 500-5000 pg/mL in 48h conditioned medium (CM) | Shifts macrophages to anti-inflammatory M2 phenotype, inhibits Th1/Th17, promotes Tregs. |
| TGF-β1 | Constitutive, enhanced by TNF-α | T cells, Macrophages | 100-1000 pg/mL in 48h CM | Suppresses T effector cell proliferation, promotes Treg differentiation. |
| IL-10 | IFN-γ, TLR agonists | Macrophages, T cells | Variable, often low (10-100 pg/mL) | Potent anti-inflammatory cytokine, deactivates macrophages. |
Protocol 1.1: Generation of MSC-Conditioned Medium (CM) for Paracrine Studies
Objective: To collect soluble factors secreted by naive MSCs under inflammatory priming. Materials:
- Human Bone Marrow-derived MSCs (Passage 3-5, characterized per ISCT criteria).
- Priming Cocktail: Recombinant human IFN-γ (50 ng/mL) + TNF-α (20 ng/mL).
- Control Medium: Serum-free, phenol-red free basal medium (e.g., DMEM/F-12).
- Equipment: CO2 incubator, centrifuge, 0.22 μm PVDF filter units.
Procedure:
- Culture MSCs to 80% confluence in T-175 flasks.
- Wash cells twice with PBS and replace medium with 20 mL of serum-free basal medium.
- Add priming cocktail to experimental flasks. Use no cytokines for control CM.
- Incubate for 48-72 hours in a standard incubator (37°C, 5% CO2).
- Collect supernatant. Centrifuge at 2000 × g for 10 min to remove cells/debris.
- Filter-sterilize using a 0.22 μm filter. Aliquot and store at -80°C. Label as "MSC-CM (Primed)" or "MSC-CM (Naive)."
Protocol 1.2: Functional T Cell Suppression Assay Using CM
Objective: To assess the functional capacity of MSC-derived soluble factors to suppress T cell proliferation. Materials:
- Peripheral Blood Mononuclear Cells (PBMCs) from healthy donor.
- CFSE Dye: For tracking cell division.
- T cell Activator: e.g., anti-CD3/CD28 coated beads.
- Flow Cytometer.
Procedure:
- Isolate PBMCs via density gradient centrifugation.
- Label PBMCs with 5 μM CFSE for 10 min at 37°C. Quench with complete medium.
- Seed CFSE-labeled PBMCs (1×10^5 per well) in a 96-well U-bottom plate.
- Add anti-CD3/CD28 beads at a 1:1 bead:cell ratio.
- Immediately add 50% (v/v) of either control medium, naive MSC-CM, or primed MSC-CM.
- Culture for 4-5 days.
- Harvest cells, stain with anti-CD3 antibody, and analyze CFSE dilution in CD3+ T cells via flow cytometry.
- Quantification: Calculate % suppression of proliferation relative to activated PBMCs in control medium.
Cell-Cell Contact: Key Receptor-Ligand Interactions and Protocols
Direct contact via adhesion molecules and receptor-ligand pairs is crucial, especially for T cell suppression. Key pathways include Programmed Death Ligand 1 (PD-L1), Vascular Cell Adhesion Molecule 1 (VCAM-1), and FAS Ligand (FASL).
Table 2: Key Contact-Dependent Mechanisms in MSCs
| MSC Surface Molecule | Receptor on Immune Cell | Inducing Signal(s) | Key Effect |
|---|---|---|---|
| PD-L1 (CD274) | PD-1 (CD279) | IFN-γ, TNF-α | Delivers inhibitory signal, inhibits T cell activation and cytokine production, promotes T cell exhaustion. |
| VCAM-1 (CD106) | VLA-4 (α4β1 integrin) | Inflammatory cytokines (TNF-α, IL-1) | Facilitates stable MSC-immune cell adhesion, necessary for localized suppressive action. |
| FASL (CD178) | FAS (CD95) | Constitutive, enhanced by priming | Induces apoptosis in activated FAS+ T cells. |
Protocol 2.1: Transwell Co-culture to Distinguish Contact vs. Paracrine Effects
Objective: To experimentally separate contact-dependent effects from paracrine-mediated immunomodulation.
Materials:
- Transwell Plates (e.g., 0.4 μm pore, 6.5 mm insert).
- Responder Immune Cells: e.g., Purified CD4+ T cells or PBMCs.
- T cell activator.
Procedure:
- Setup Conditions:
- A. Contact: Seed MSCs in the lower well. Add responder T cells directly onto MSCs.
- B. Paracrine (Transwell): Seed MSCs in the lower well. Place responder T cells in the insert (upper chamber).
- C. Control: T cells alone in lower well with/without activator.
- Prime MSCs in all relevant wells with IFN-γ (50 ng/mL) for 24h prior to adding immune cells.
- Activate T cells in all conditions with soluble anti-CD3/CD28.
- Co-culture for 72 hours.
- Harvest T cells from each condition and assess:
- Proliferation: Via CFSE dilution or 3H-thymidine incorporation.
- Cytokine Profile: ELISA for IFN-γ (Th1) and IL-17 (Th17) in supernatant.
- Apoptosis: Annexin V/PI staining by flow cytometry.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Core Mechanism Analysis
| Reagent / Kit | Provider Examples | Function in Experimental Design |
|---|---|---|
| Recombinant Human IFN-γ | PeproTech, R&D Systems | Gold-standard cytokine to prime/license MSCs for immunomodulation. |
| IDO Activity Assay Kit | Sigma-Aldrich, Abcam | Measures kynurenine production to quantify functional IDO enzyme activity in MSC cultures. |
| PGE2 Parameter Assay Kit | R&D Systems, Cayman Chemical | Quantifies PGE2 concentration in MSC-CM via ELISA. |
| Human TGF-β1 ELISA Kit | BioLegend, Thermo Fisher | Measures latent and active TGF-β1 levels in CM. |
| Anti-Human PD-L1 Blocking Antibody | BioLegend, eBioscience | Used in neutralization experiments to disrupt PD-1/PD-L1 contact pathway. |
| CFSE Cell Division Tracker | Thermo Fisher, BioLegend | Fluorescent dye to track and quantify lymphocyte proliferation. |
| Annexin V Apoptosis Detection Kit | BD Biosciences | To assess contact-dependent induction of apoptosis in T cells. |
| Transwell Permeable Supports | Corning | Crucial for separating contact and paracrine effects in co-culture. |
Visualizations
Title: MSC Immunomodulation Core Mechanism Flow
Title: Transwell Assay Design for Paracrine Effects
Within the thesis on Genetic modification of MSCs to enhance immunomodulation, understanding and leveraging key soluble factors is paramount. Mesenchymal stromal cells (MSCs) mediate immunosuppression primarily through paracrine secretion, with Indoleamine 2,3-dioxygenase (IDO), Prostaglandin E2 (PGE2), Transforming Growth Factor-beta (TGF-β), and Interleukin-10 (IL-10) being critical players. These factors collectively modulate dendritic cell maturation, drive T regulatory cell (Treg) expansion, suppress pro-inflammatory T helper 1 (Th1) and T helper 17 (Th17) responses, and inhibit cytotoxic T cell and NK cell function. Genetic engineering strategies aim to overexpress these factors or the enzymes that produce them (e.g., IDO1, PTGES2 for PGE2 synthesis) to create "super-immunosuppressive" MSCs with enhanced potency and persistence for treating autoimmune diseases, graft-versus-host disease (GvHD), and allograft rejection.
Table 1: Key Immunomodulatory Soluble Factors
| Factor | Primary Cellular Source | Key Immunological Targets | Primary Effect | Typical Concentration Range in MSC-Conditioned Media* |
|---|---|---|---|---|
| IDO | MSCs (IFN-γ-induced), DCs | T cells, NK cells | Depletes tryptophan, increases kynurenines; inhibits T/NK proliferation, induces Tregs. | Enzyme activity: 20-50 µM Kynurenine/10^6 cells/24h. |
| PGE2 | MSCs (constitutive & induced) | Macrophages (M1→M2), DCs, Th1/Th17, Tregs | Shifts macrophages to M2, inhibits DC maturation, suppresses Th1/Th17, promotes Tregs. | 1-10 ng/ml per 10^6 cells/24h. |
| TGF-β | MSCs, Tregs, many stromal cells | CD4+ T cells, CD8+ T cells, B cells, Macrophages | Potent suppression of T-cell proliferation & effector functions; promotes Treg differentiation. | 100-500 pg/ml per 10^6 cells/24h. |
| IL-10 | MSCs (often induced), Tregs, M2 Macrophages | Antigen-Presenting Cells (APCs), T cells | Deactivates APCs, inhibits pro-inflammatory cytokine production, enhances B cell survival. | 50-200 pg/ml per 10^6 cells/24h (highly variable). |
Note: Concentrations are highly dependent on MSC source, passage, culture conditions, and inflammatory priming (e.g., with IFN-γ/TNF-α).
Table 2: Genetic Modification Strategies to Enhance Factor Secretion
| Target Factor | Gene(s) for Overexpression | Common Vector | Expected Enhancement Fold (Range) | Primary Therapeutic Target Model |
|---|---|---|---|---|
| IDO | IDO1 | Lentivirus, Adenovirus | 5-20x enzyme activity | GvHD, Autoimmune Encephalomyelitis |
| PGE2 | PTGES1 (COX-2/mPGES-1) | Lentivirus, Plasmid | 3-10x PGE2 secretion | Colitis, Acute Lung Injury |
| TGF-β | TGFB1 (active form) | Lentivirus | 10-50x protein secretion | Cardiac Allograft, Renal Fibrosis |
| IL-10 | IL10 | Lentivirus, AAV | 50-100x protein secretion | Colitis, Arthritis, Ischemia-Reperfusion |
Experimental Protocols
Protocol 3.1: In Vitro T-Cell Suppression Assay Using Genetically Modified MSCs
Objective: To quantify the enhanced immunosuppressive capacity of IDO- or PGE2-overexpressing MSCs via suppression of T-cell proliferation. Materials: Human PBMCs, CFSE, anti-CD3/CD28 beads, IDO1-overexpressing MSCs (IDO-MSCs), control MSCs, IFN-γ, 1-Methyl-DL-tryptophan (1-MT, IDO inhibitor), PGE2 receptor antagonist (e.g., AH6809). Procedure:
- Isolate PBMCs from healthy donor blood via density gradient centrifugation.
- Label PBMCs with 5 µM CFSE for 10 min at 37°C. Quench with complete RPMI-1640.
- Plate 1x10^5 CFSE-labeled PBMCs per well in a 96-well U-bottom plate. Activate T cells with anti-CD3/CD28 beads (bead:cell ratio 1:1).
- In transwell inserts or in direct contact, add either wild-type (WT) MSCs or genetically modified MSCs (IDO-MSCs) at MSC:PBMC ratios of 1:10, 1:20, and 1:50.
- For inhibition controls, add 500 µM 1-MT (for IDO blockade) or 10 µM AH6809 (for PGE2 blockade) to relevant wells.
- Culture for 4-5 days. Harvest cells and analyze CFSE dilution of CD3+ T cells by flow cytometry to determine proliferation index.
- Calculate % Suppression = [1 - (Proliferation with MSCs / Proliferation without MSCs)] * 100.
Protocol 3.2: Quantification of Soluble Factors via ELISA
Objective: To validate the enhanced secretory profile of genetically modified MSCs post-inflammatory priming. Materials: WT and modified MSCs, IFN-γ (10 ng/ml) and TNF-α (5 ng/ml), serum-free media, ELISA kits for human IDO (via Kynurenine), PGE2, TGF-β1 (latent & active), IL-10. Procedure:
- Seed MSCs at 2x10^4 cells/well in a 24-well plate. Allow to adhere overnight.
- Replace medium with 500 µl serum-free medium +/- IFN-γ/TNF-α cytokine mix.
- Condition for 48 hours. Collect supernatant and centrifuge at 3000xg to remove debris.
- For IDO activity: Mix supernatant with 30% trichloroacetic acid, heat, and centrifuge. Add supernatant to Ehrlich's reagent. Measure absorbance at 490nm. Compare to a kynurenine standard curve.
- For PGE2, TGF-β1 (must acid-activate latent form), and IL-10: Perform assays according to commercial ELISA kit instructions using undiluted or appropriately diluted supernatant.
- Normalize concentrations to total cellular protein (via BCA assay) or cell count.
Protocol 3.3: In Vivo Assessment in a GvHD Model
Objective: To evaluate the therapeutic efficacy of TGF-β/IL-10 co-expressing MSCs in a xenogeneic GvHD mouse model. Materials: NOD-scid IL2Rγnull (NSG) mice, human PBMCs (for xenogeneic GvHD), luciferase-expressing TGF-β/IL-10-MSCs, IVIS imaging system, clinical GvHD scoring system. Procedure:
- Induce GvHD in NSG mice by intravenous (IV) injection of 5x10^5 human PBMCs (Day 0).
- On Day +1 and Day +7, administer 1x10^6 luciferase-tagged TGF-β/IL-10-MSCs or WT-MSCs via IV injection.
- Monitor mice daily for weight loss, posture, activity, fur texture, and skin integrity. Assign a clinical score (0-10).
- Weekly, inject luciferin substrate and image mice using IVIS to track MSC biodistribution.
- At endpoints (e.g., Day 28 or severe morbidity), analyze serum for human cytokine levels (ELISA) and harvest spleen/liver for histopathology and flow cytometry to assess human T cell infiltration and phenotype (Tregs vs. effector T cells).
Signaling Pathways & Workflow Diagrams
Title: IDO-Mediated Immunomodulation Pathway
Title: PGE2, TGF-β, and IL-10 Synergistic Actions
Title: Key Experimental Workflow for Engineered MSCs
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for MSC Immunomodulation Research
| Reagent Category | Specific Item/Kit | Primary Function in Research |
|---|---|---|
| MSC Characterization | Human MSC Analysis Kit (BD) | Flow cytometry-based verification of ISCT-defined MSC surface markers (CD73+, CD90+, CD105+, CD45-). |
| Inflammatory Priming | Recombinant Human IFN-γ & TNF-α | To mimic inflammatory milieu and induce maximal expression of endogenous immunomodulatory factors (IDO, PGE2). |
| Genetic Modification | Lentiviral ORF Clones (IDO1, TGFB1, IL10) | For stable, high-efficiency overexpression of target genes in MSCs. |
| Factor Quantification | PGE2 Parameter Assay Kit (R&D), Human TGF-β1 ELISA (BioLegend) | Accurate measurement of soluble factor concentrations in conditioned media or serum. |
| IDO Activity Assay | IDO Activity Assay Kit (Sigma) | Colorimetric measurement of kynurenine production to assess functional IDO enzyme levels. |
| T-cell Suppression CFSE Cell Division Tracker Kit (BioLegend), anti-CD3/CD28 Activator Beads | To label and activate T cells for co-culture suppression assays with MSCs. | |
| Flow Cytometry Antibodies | Anti-human CD4, CD25, FoxP3 (for Tregs); CD206 (for M2 macrophages) | To analyze immune cell phenotype shifts induced by engineered MSC secretome. |
| Pathway Inhibitors | 1-Methyl-DL-tryptophan (1-MT, IDO inhibitor), AH6809 (PGE2 receptor antagonist), SB431542 (TGF-β receptor inhibitor) | To confirm the specific mechanistic role of each factor in functional assays. |
| In Vivo Imaging | Luciferase-expressing Lentivirus, D-Luciferin | To track the biodistribution and persistence of administered engineered MSCs in live animals. |
Application Notes: Quantifying Variability in MSC Immunomodulation
The clinical translation of mesenchymal stromal cells (MSCs) for immunomodulatory therapies is significantly hampered by donor-to-donor and batch-to-batch variability, leading to inconsistent therapeutic outcomes—a "Crisis of Effect." Genetic modification presents a strategy to standardize and enhance potency. The following data, compiled from recent studies, quantifies this variability and the impact of key genetic modifications.
Table 1: Sources of Variability in Native MSC Immunomodulatory Potency
| Variability Source | Measured Parameter | Range/Fold-Difference | Impact on Clinical Outcome (Example) |
|---|---|---|---|
| Donor Heterogeneity | PGE2 Secretion (Baseline) | Up to 100-fold between donors | Graft-vs-Host-Disease (GVHD) response unpredictable |
| Passage Number | IDO1 Activity (After IFN-γ stimulation) | ~80% decrease from P3 to P8 | Late-passage cells fail to suppress T-cell proliferation |
| Serum Lot | TSG-6 Expression | 3 to 5-fold variation | Efficacy in acute lung injury models inconsistent |
| Cell Density at Harvest | COX-2 mRNA Levels | 2 to 4-fold variation | Altered prostaglandin-mediated macrophage polarization |
Table 2: Efficacy of Genetic Modifications to Enhance & Standardize Immunomodulation
| Genetic Modification (Target) | Model System | Quantified Enhancement vs. Naive MSCs | Key Standardization Metric |
|---|---|---|---|
| Overexpression of IDO1 | Human PBMC Co-culture | T-cell proliferation inhibited by 95% ± 3% (vs. 60% ± 25% in naive) | Coefficient of variation (CV) of suppression reduced from 42% to 8% |
| Forced Expression of PD-L1 | Mouse Colitis Model | Clinical score improvement: 70% vs. 40% | Serum IL-6 levels: 150 pg/mL ± 20 pg/mL (modified) vs. 300 ± 150 pg/mL (naive) |
| CRISPRa activation of TSG-6 | LPS-induced Macrophage Co-culture | TNF-α suppression: 90% ± 5% (vs. 50% ± 30%) | Dynamic range of TNF-α suppression narrowed by 75% |
| Knock-in of Inducible IL-10 | Mouse Skin Allograft | Graft survival extended to 28 ± 2 days (vs. 16 ± 7 days) | Inter-animal survival time standard deviation reduced 3-fold |
Detailed Experimental Protocols
Protocol 1: StandardizedIn VitroPotency Assay for Genetically Modified MSCs
Purpose: To quantitatively compare the immunomodulatory capacity of engineered MSCs against donor-matched naive MSCs using a T-cell proliferation assay. Key Reagents: See "Scientist's Toolkit" (Table 3). Procedure:
- MSC Preparation: Seed donor-matched naive and genetically modified (e.g., IDO1-overexpressing) MSCs at 2.5 x 10^4 cells/well in a 48-well plate. Culture for 24 hours.
- PBMC Activation: Isolate CD3+ T-cells from human PBMCs using negative selection. Stain with CFSE (5 µM) per manufacturer's protocol.
- Co-culture: Add 1 x 10^5 CFSE-stained T-cells to MSC monolayers. Activate T-cells with soluble anti-CD3/CD28 antibodies (1 µg/mL each). Include MSC-only and T-cell-only controls.
- Incubation: Culture for 96 hours in a 37°C, 5% CO2 incubator.
- Analysis: Harvest non-adherent cells. Analyze CFSE dilution via flow cytometry. Calculate percentage suppression of proliferation:
[1 - (%Dividing T-cells in co-culture / %Dividing T-cells in T-cell-only control)] x 100. - Statistical Standardization: Calculate the Coefficient of Variation (CV) across n≥3 biological replicates for each MSC type. A lower CV indicates higher standardization.
Protocol 2:In VivoValidation in a Mouse Model of Graft-vs-Host-Disease (GVHD)
Purpose: To evaluate the consistency of therapeutic effect of PD-L1-overexpressing MSCs. Key Reagents: See "Scientist's Toolkit" (Table 3). Procedure:
- GVHD Induction: Irradiate (8.5 Gy) recipient BALB/c mice. Within 24 hours, inject 5 x 10^6 bone marrow cells and 2 x 10^6 splenocytes from C57BL/6 donors via tail vein.
- Therapy Administration: On day +1 and +4, administer 1 x 10^6 naive or PD-L1-overexpressing human MSCs (in PBS) via intravenous injection. Include a PBS-treated control group (n=10 per group).
- Clinical Scoring: Monitor mice daily from day +5. Score for five clinical parameters (weight loss, posture, activity, fur texture, skin integrity) on a scale of 0-2 each. Sum for a maximum score of 10.
- Endpoint Analysis: Record survival daily. At a pre-defined endpoint (e.g., day +28), sacrifice and collect serum and target organs (skin, liver, intestine).
- Consistency Metric: Measure serum IFN-γ by ELISA. Compare not only the mean level between groups but also the standard deviation and range as measures of effect consistency.
Visualization: Pathways and Workflows
Diagram Title: Engineering Strategy to Overcome MSC Variability
Diagram Title: Potency Validation Workflow for Engineered MSCs
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions for MSC Potency Engineering & Assays
| Reagent / Material | Function & Relevance | Example Product/Catalog |
|---|---|---|
| Lentiviral IDO1 Overexpression Vector | Drives consistent, high-level expression of indoleamine 2,3-dioxygenase, a key immunomodulatory enzyme, overcoming donor variability. | VectorBuilder, VB010000-1342ztt |
| Recombinant Human IFN-γ | Critical stimulus to induce immunomodulatory pathways in MSCs; used to test inducible gene circuits and validate functionality. | PeproTech, 300-02 |
| Anti-human CD3/CD28 Activator | Activates T-cells in co-culture potency assays to measure MSC-mediated suppression of proliferation. | Stemcell Technologies, 10971 |
| CFSE Cell Division Tracker | Fluorescent dye to quantitatively track T-cell proliferation in co-culture assays via flow cytometry. | Thermo Fisher, C34554 |
| IDO Activity Assay Kit | Quantifies kynurenine production, providing a direct biochemical readout of IDO1 enzymatic function in modified MSCs. | Sigma-Aldrich, MAK315 |
| Mouse GvHD Induction Kit | Standardized bone marrow and spleen cells for reliable, consistent induction of GvHD in murine efficacy studies. | Jackson Labs, Stock 100013 |
| Multiplex Cytokine ELISA Panel | Simultaneously measures key inflammatory (IFN-γ, TNF-α, IL-6) and regulatory (IL-10) cytokines from serum or supernatant. | R&D Systems, LXSAHM |
| CRISPRa sgRNA for TSG-6 | Guide RNA for CRISPR activation to upregulate endogenous TSG-6 expression without transgene integration. | Synthego, custom design |
Application Notes
The strategic genetic modification of Mesenchymal Stromal Cells (MSCs) for enhanced immunomodulation requires precise targeting of key regulatory pathways. The rationale is based on overcoming the inherent limitations of naïve MSCs, such as variable potency, transient therapeutic effects, and suboptimal homing. By enhancing specific signaling nodes and secretory profiles, engineered MSCs can achieve consistent, potent, and durable immunomodulatory outcomes for treating autoimmune diseases, graft-versus-host disease (GvHD), and inflammatory disorders.
Recent data highlights the most promising targets:
Table 1: Key Immunomodulatory Pathways for MSC Genetic Enhancement
| Target Pathway/Gene | Primary Function | Quantitative Enhancement Post-Modification | Key Effect on Immune Response |
|---|---|---|---|
| IDO1 (Indoleamine 2,3-dioxygenase 1) | Tryptophan catabolism to kynurenines | Enzyme activity increased 3-5 fold vs. naïve MSC. Kynurenine levels in co-culture: 50-70 µM vs. 15-25 µM (naïve). | Robust suppression of T-cell proliferation (70-90% inhibition). Promotes Treg differentiation. |
| PD-L1 (Programmed Death-Ligand 1) | Immune checkpoint engagement | Surface expression increased 8-12 fold (MFI). | Enhances apoptosis of activated T-cells. Synergizes with IDO1. |
| TSG-6 (TNF-α Stimulated Gene 6) | Anti-inflammatory, ECM modulator | Secretion elevated >10 ng/10^6 cells/24h vs. ~1 ng in naïve. | Potent inhibition of neutrophil infiltration, NF-κB signaling. Critical in sepsis/ARDS models. |
| IL-10 (Interleukin-10) | Master anti-inflammatory cytokine | Secretion to 500-1000 pg/mL/24h vs. often undetectable baseline. | Polarizes macrophages to M2 phenotype, reduces Th1/Th17 cytokines. |
| HGF (Hepatocyte Growth Factor) | Paracrine regenerative & immunomodulatory factor | Secretion increased 4-6 fold (~8-12 ng/mL/24h). | Promotes Treg function, inhibits dendritic cell maturation. |
| FOXP3 | Master regulator of Tregs | Ectopic expression directs MSC to acquire Treg-like suppressor function. | Enables contact-independent suppression, stable phenotype. |
Justification for Target Selection:
- IDO1 & PD-L1: Constitute a combinatorial "immunological synapse" for T-cell regulation. Essential for overcoming the inflammatory cytokine storm that can deactivate therapeutic MSCs.
- TSG-6: A critical mediator for sterile inflammatory conditions. Its enhancement protects MSCs and provides potent bystander anti-inflammatory effects.
- Paracrine Factor Cocktail (IL-10, HGF): Moves beyond single-factor therapy. Genetically programming a sustained, multi-factorial secretory profile mimics the peak potency of activated MSCs constitutively.
- FOXP3: Repurposes MSC transcriptional identity, creating a novel stromal-immunoregulatory hybrid cell with potentially greater stability and specificity.
Experimental Protocols
Protocol 1: Lentiviral Transduction for Overexpression of IDO1 and PD-L1 in Human MSCs
Objective: To generate genetically enhanced human MSCs (heMSCs) stably overexpressing IDO1 and PD-L1.
Materials (Research Reagent Solutions):
- Human Bone Marrow-derived MSCs: (P3-P5), validated for trilineage differentiation and surface markers (CD73+, CD90+, CD105+, CD45-).
- Lentiviral Vectors: Third-generation, VSV-G pseudotyped, encoding human IDO1 and PD-L1 genes under EF1α promoter, with fluorescent (e.g., GFP) or puromycin selection markers.
- Polybrene (Hexadimethrine bromide): Enhances viral transduction efficiency.
- Puromycin Dihydrochloride: For selection of stably transduced cells.
- Complete MSC Medium: α-MEM, 10% FBS (qualified for MSCs), 1% GlutaMAX.
- Flow Cytometry Antibodies: Anti-human CD274 (PD-L1) PE, Isotype control.
Procedure:
- Day 0: Seeding. Plate MSCs at 8,000 cells/cm² in complete medium.
- Day 1: Transduction. Replace medium with fresh medium containing 8 µg/mL Polybrene. Add lentiviral particles at a predetermined Multiplicity of Infection (MOI of 10-20). Include a mock-transduction control (virus-free).
- Day 2: Medium Change. Replace with fresh complete medium.
- Day 4: Selection. Begin selection with 1-2 µg/mL puromycin. Maintain selection for 7 days, changing medium every 2-3 days.
- Day 12: Validation.
- Flow Cytometry: Harvest cells, stain for PD-L1 surface expression. Analyze fold-change in Mean Fluorescence Intensity (MFI) vs. mock control.
- Functional Assay (IDO1): Seed heMSCs at 50,000 cells/well in 24-well plate. Stimulate with 100 ng/mL IFN-γ for 48h. Collect supernatant and measure kynurenine concentration via spectrophotometric assay (absorbance 490nm) after reaction with Ehrlich's reagent. Compare to mock-transduced, IFN-γ-stimulated MSCs.
Protocol 2: In Vitro T-cell Suppression Assay with heMSCs
Objective: To quantify the enhanced immunomodulatory capacity of IDO1/PD-L1 heMSCs on activated human T-cell proliferation.
Materials:
- Responder T-cells: Isolated PBMCs from healthy donor, or purified CD3+ T-cells.
- Activation Beads: Anti-human CD3/CD28 Dynabeads.
- CFSE (Carboxyfluorescein succinimidyl ester): For tracking T-cell division.
- Co-culture Plates: 96-well round-bottom plates.
- Flow Cytometry Buffer: PBS + 2% FBS.
Procedure:
- T-cell Labeling: Isolate CD3+ T-cells. Resuspend at 1-2x10^6/mL in PBS/0.1% BSA. Add CFSE to final 0.5-1 µM, incubate 10 min at 37°C. Quench with 5x volume of cold complete RPMI. Wash twice.
- Co-culture Setup: Plate heMSCs or control MSCs (irradiated with 20 Gy to prevent proliferation) in 96-well plate at ratios of 1:10, 1:20, and 1:50 (MSC:T-cell). Add 100,000 CFSE-labeled T-cells per well. Activate T-cells with CD3/CD28 beads (bead:cell ratio 1:1). Set up T-cell only (max proliferation) and unstimulated T-cell (background) controls. Use 200 µL total volume/well.
- Culture: Incubate for 4-5 days in a humidified 37°C, 5% CO2 incubator.
- Analysis: Harvest cells, acquire on flow cytometer using FL1 channel for CFSE. Gate on live lymphocyte population. Calculate percentage of proliferated T-cells (CFSE-low) and division index. Calculate % Suppression = [1 - (% Proliferation in Co-culture / % Proliferation in T-cell only control)] x 100.
Visualizations
Diagram 1: IDO1 & PD-L1 synergistic immunomodulation pathway.
Diagram 2: Workflow for generating and testing genetically enhanced MSCs.
The Scientist's Toolkit
Table 2: Essential Research Reagents for MSC Immunomodulation Engineering
| Reagent/Material | Function in Protocol | Key Consideration |
|---|---|---|
| Lentiviral Vectors (3rd Gen) | Safe, efficient delivery and stable integration of target genes into MSC genome. | Use high-titer (>1x10^8 IU/mL), functional titer on MSCs must be pre-determined. |
| Polybrene | Cationic polymer that neutralizes charge repulsion, increasing virus-cell contact. | Optimize concentration (4-10 µg/mL); can be cytotoxic at high levels or with prolonged exposure. |
| Puromycin Dihydrochloride | Selects for stably transduced cells by eliminating non-transduced population. | Critical to determine kill curve (typical range 0.5-2 µg/mL for MSCs) before experiment. |
| Recombinant Human IFN-γ | Gold-standard inflammatory cytokine to stimulate endogenous immunomodulatory pathways in MSCs. | Used for functional validation (e.g., IDO1 assay). High purity, carrier-free recommended. |
| CFSE Cell Division Dye | Fluorescent dye that dilutes 2-fold with each cell division, allowing precise quantification of T-cell proliferation suppression. | Aliquot and store at -20°C protected from light. Optimize labeling concentration for target cell type. |
| Anti-human CD3/CD28 Activator Beads | Provides strong, reproducible, and soluble-free activation of T-cells for suppression assays. | Bead-to-cell ratio is crucial; 1:1 is standard but may require titration. |
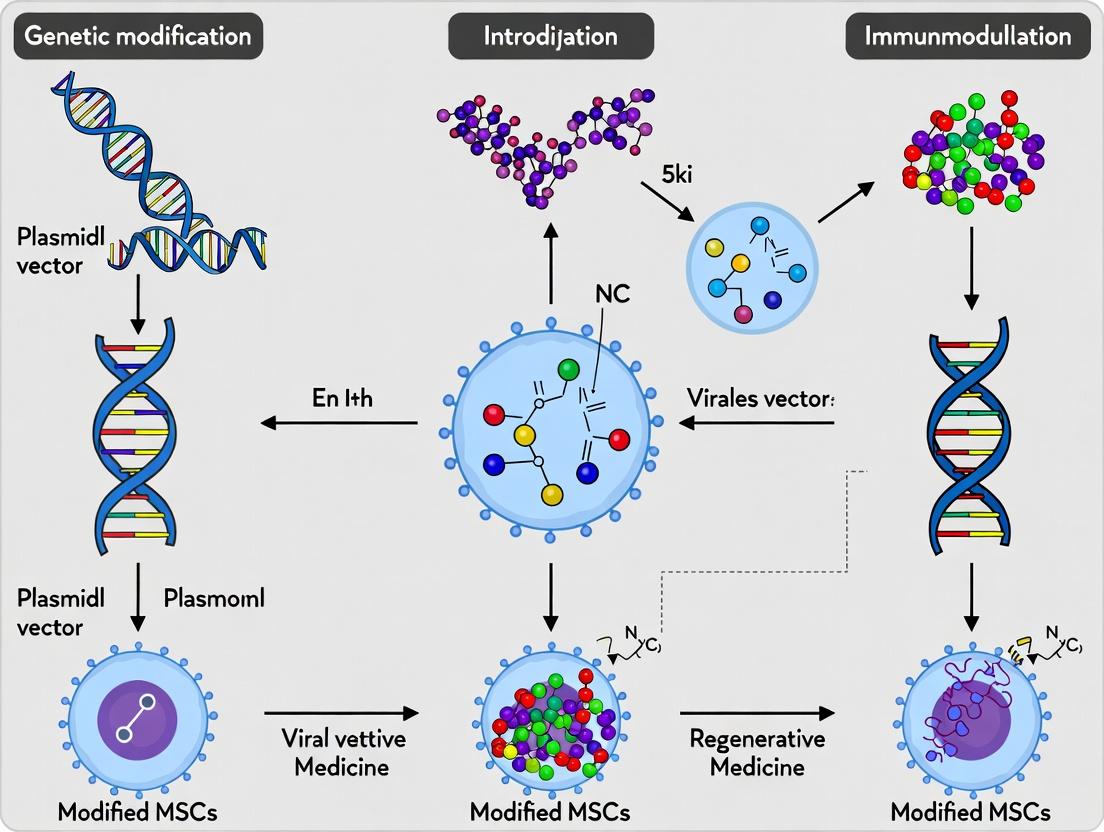
Building a Better MSC: Cutting-Edge Gene Delivery and Editing Techniques
Within the context of a broader thesis on the genetic modification of mesenchymal stromal cells (MSCs) to enhance their immunomodulatory potential, the selection of an appropriate viral vector is a foundational decision. Lentiviral and adenoviral vectors are two predominant workhorses, each offering distinct advantages for achieving stable genomic integration or high-level transient expression, respectively. This application note details their comparative virology, protocols for MSC transduction, and key considerations for immunomodulation research.
Comparative Vector Characteristics
Table 1: Core Characteristics of Lentiviral vs. Adenoviral Vectors for MSC Modification
| Parameter | Lentivirus (e.g., HIV-1 based) | Adenovirus (e.g., Ad5 based) |
|---|---|---|
| Nucleic Acid | Single-stranded RNA | Double-stranded DNA |
| Integration | Stable (integrates into host genome) | Transient (episomal) |
| Therapeutic Expression Duration | Long-term, sustained (weeks-months) | Short-term, high-level (days-weeks) |
| Typimal Titer (Functional) | 1 x 10^8 - 1 x 10^9 TU/mL | 1 x 10^10 - 1 x 10^12 VP/mL |
| Transduction of Dividing/Non-dividing Cells | Efficient for both | Requires cell division for nuclear entry; less efficient in quiescent cells |
| Typical Transduction Efficiency in MSCs | 30-80% (dose-dependent) | 70-95% (dose-dependent) |
| Packaging Capacity | ~8 kb | ~7.5 kb (first-generation) / ~36 kb (helper-dependent) |
| Immunogenicity | Lower in vitro | High; triggers strong innate & adaptive immune responses |
| Common Biosafety Level | BSL-2+ | BSL-2 |
Table 2: Application Selection for MSC Immunomodulation Research
| Research Goal | Recommended Vector | Rationale |
|---|---|---|
| Long-term in vivo tracking or secretome studies | Lentivirus | Stable integration ensures transgene persistence through cell divisions. |
| Overexpression of a therapeutic factor (e.g., IL-10, IDO) for chronic disease models | Lentivirus | Sustained, consistent expression is required for long-term effects. |
| Rapid screening of multiple candidate genes in vitro | Adenovirus | High-titer, rapid expression allows quick functional assays. |
| Transient, potent immunomodulation in acute inflammation models | Adenovirus | High-level, burst expression can powerfully modulate acute immune responses. |
| Knockdown studies (shRNA) | Lentivirus | Required for stable, long-term gene silencing. |
| Expression of large or multiple transgenes | High-Capacity Adenovirus | Accommodates large genetic payloads without integration. |
Detailed Protocols
Protocol 1: Lentiviral Transduction of Human Bone Marrow-Derived MSCs for Stable Expression
Objective: To generate a polyclonal population of MSCs stably expressing an immunomodulatory transgene (e.g., TGF-β).
Materials (Research Reagent Solutions):
- Polybrene (Hexadimethrine bromide): A cationic polymer that enhances viral adhesion to the cell membrane.
- Lentiviral particles: Third-generation, VSV-G pseudotyped, encoding the gene of interest (GOI) under a constitutive promoter (e.g., EF1α, PGK).
- MSC Growth Medium: α-MEM supplemented with 10% FBS, 1% GlutaMAX, 1% Penicillin/Streptomycin.
- Puromycin or appropriate antibiotic: For selection of transduced cells post-transduction.
Method:
- Day 0: Seeding. Plate early-passage (P3-P5) human MSCs at 50-70% confluence (approx. 1 x 10^4 cells/cm²) in standard growth medium.
- Day 1: Transduction. a. Pre-warm fresh growth medium and viral aliquots. b. Prepare the transduction mix: Growth medium containing the calculated lentiviral particles (MOI of 5-20 is typical for MSCs) and 8 µg/mL Polybrene. Mix gently. c. Aspirate medium from cells and add the transduction mix. d. Incubate cells at 37°C, 5% CO₂ for 16-24 hours.
- Day 2: Medium Change. Carefully aspirate the transduction mix and replace with fresh, pre-warmed growth medium.
- Days 3-5: Selection. Begin antibiotic selection (e.g., 1-2 µg/mL puromycin) 48-72 hours post-transduction. Maintain selection for 5-7 days, changing medium every 2-3 days, until all non-transduced control cells are dead.
- Validation: Expand the polyclonal population and validate transgene expression via flow cytometry (for surface markers) or Western blot/ELISA (for secreted factors).
Protocol 2: Adenoviral Transduction of MSCs for High-Level Transient Expression
Objective: To achieve high, transient overexpression of a cytokine (e.g., IL-10) in MSCs for an acute in vitro co-culture assay.
Materials (Research Reagent Solutions):
- Adenoviral particles: First-generation, E1/E3-deleted, expressing the GOI under a strong promoter (e.g., CMV).
- Phosphate-Buffered Saline (PBS), Mg²⁺/Ca²⁺-free: For washing cells.
- MSC Serum-Free Medium or Reduced-Serum Medium: Used during transduction to improve viral adsorption.
Method:
- Day 0: Seeding. Plate MSCs at 60-80% confluence to ensure they are actively dividing.
- Day 1: Transduction. a. Wash cells once with pre-warmed PBS. b. Prepare the viral inoculum in serum-free or reduced-serum (2% FBS) medium. The required MOI for MSCs is typically higher than for cell lines, often ranging from 100-1000 particle units (PU) per cell. c. Add the inoculum to the cells. d. Incubate at 37°C, 5% CO₂ for 1-2 hours, gently rocking the plate every 15-20 minutes. e. After incubation, add an equal volume of complete growth medium (containing 20% FBS) to achieve a final serum concentration of ~10%. Alternatively, replace the inoculum with fresh complete medium. f. Return cells to the incubator.
- Day 2-4: Assay. Peak transgene expression is typically observed 24-72 hours post-transduction. Harvest conditioned medium for cytokine analysis or use the transduced MSCs in functional immunomodulation assays (e.g., T-cell suppression assays).
Signaling & Workflow Visualizations
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Viral Transduction of MSCs
| Reagent | Function & Importance in MSC Context | Typical Vendor/Example |
|---|---|---|
| VSV-G Pseudotyped Lentivirus | Broad tropism allows efficient transduction of hard-to-transfect MSCs. Essential for stable gene delivery. | Prepared in-house via 3rd-gen packaging system (psPAX2, pMD2.G) or from core facilities. |
| Hexadimethrine bromide (Polybrene) | Neutralizes charge repulsion between virus and cell membrane, enhancing transduction efficiency. Critical for low-MOI lentiviral work. | Sigma-Aldrich, Millipore. |
| Puromycin Dihydrochloride | Selection antibiotic for lentiviral constructs containing a puromycin resistance gene (PAC). Allows rapid enrichment of transduced MSC populations. | Thermo Fisher, Invivogen. |
| First-Generation Adenovirus (E1/E3 deleted) | Provides high transduction efficiency and rapid, high-level transgene expression in MSCs for transient studies. | Vector Biolabs, Vigene Biosciences. |
| Cell Recovery Solution (or Detachin) | Gentle, enzymatic-free dissociation reagent. Preserves MSC surface markers and viability post-transduction better than trypsin. | Corning, ScienCell. |
| MSC-Qualified Fetal Bovine Serum (FBS) | Supports robust MSC growth and maintains multipotency. Batch testing is crucial to ensure consistent viral transduction and cell health. | Gibco, Thermo Fisher. |
| Lentivirus Titer Kit (qPCR-based) | Accurately quantifies functional viral titer (TU/mL) for calculating precise MOI, critical for reproducible experiments. | Takara Bio, Abcam. |
| Human MSC Phenotyping Kit (Flow Cytometry) | Validates MSC identity (CD73+, CD90+, CD105+, CD34-, CD45-) post-transduction/manipulation. | Miltenyi Biotec, BD Biosciences. |
Within the central thesis of enhancing mesenchymal stem cell (MSC) immunomodulatory function through genetic modification, the selection of a safe, efficient, and persistent transfection strategy is paramount. Viral vectors, while efficient, pose safety risks such as insertional mutagenesis and immunogenicity, complicating clinical translation. This application note details three leading non-viral strategies—electroporation, lipofection, and transposon systems—providing comparative data, optimized protocols for MSCs, and essential toolkit resources for researchers.
Electroporation (Nucleofection) for MSCs
Electroporation uses electrical pulses to create transient pores in the cell membrane, allowing direct cytoplasmic delivery of nucleic acids. "Nucleofection," a specialized form, targets the nucleus for higher efficiency in hard-to-transfect primary cells like MSCs.
Key Application: Rapid, high-efficiency delivery of plasmid DNA, mRNA, or CRISPR-Cas9 ribonucleoprotein (RNP) complexes for in vitro functional screening of immunomodulatory genes (e.g., overexpression of IDO, IL-10, or PD-L1).
Protocol: Nucleofection of Human Bone Marrow-Derived MSCs with Plasmid DNA
- Cell Preparation: Culture passage 3-5 MSCs in α-MEM + 10% FBS. Harvest at 80-90% confluence using trypsin/EDTA. Count and pellet 1 x 10^6 cells.
- DNA Complexation: Resuspend cell pellet in 100 µL of Room Temperature Human MSC Nucleofector Solution (Lonza, VPE-1001). Add 2-5 µg of endotoxin-free plasmid DNA (e.g., pMAX-GFP for optimization). Mix gently.
- Nucleofection: Transfer cell/DNA mixture to a certified cuvette. Run the pre-optimized program for human MSCs (e.g., U-23 on Lonza 4D-Nucleofector). Immediately add 500 µL of pre-warmed culture medium to the cuvette.
- Recovery & Analysis: Transfer cells to a 6-well plate with 2 mL complete medium. Assess transfection efficiency via flow cytometry (for reporter genes) at 24-48 hours. For immunomodulation assays, perform functional analysis (e.g., T-cell suppression assay) 72+ hours post-transfection.
Quantitative Data Summary: Electroporation Table 1: Performance Metrics of Electroporation in MSCs
| Parameter | Typical Range | Notes |
|---|---|---|
| Transfection Efficiency | 40-80% | Highly dependent on MSC source, passage, and optimization. |
| Cell Viability (24h post) | 60-75% | Lower viability is common; requires careful recovery. |
| Max Cargo Size | >10 kbp (plasmid) | Suitable for large plasmids. Also effective for mRNA & RNP. |
| Onset of Expression | Rapid (hours) | mRNA/RNP: hours; plasmid DNA: 12-24 hours. |
| Transgene Duration | Transient (days-weeks) | Episomal expression; diluted with cell division. |
Diagram 1: Electroporation workflow for MSC transfection.
Lipofection (Lipid Nanoparticle-Mediated Transfection)
Lipofection utilizes cationic lipids that self-assemble with nucleic acids to form lipoplexes, which fuse with the cell membrane and release cargo into the cytoplasm.
Key Application: Simple, high-throughput screening of siRNA, miRNA, or plasmid DNA to knock down or overexpress genes modulating MSC paracrine signaling pathways.
Protocol: Lipofection of Adipose-Derived MSCs with siRNA
- Cell Seeding: Seed 1.5 x 10^5 MSCs/well in a 12-well plate 24 hours prior to achieve 60-70% confluence at transfection.
- Complex Preparation: Dilute 5 µL of a 20 µM siRNA stock (against target, e.g., STAT3) in 100 µL serum-free Opt-MEM (Tube A). Dilute 5 µL of a commercial lipid reagent (e.g., Lipofectamine RNAiMAX) in 100 µL Opt-MEM (Tube B). Incubate 5 min at RT.
- Lipoplex Formation: Combine Tube A and Tube B. Mix gently and incubate 20 min at RT.
- Transfection: Add 200 µL of lipoplex mixture dropwise to each well containing 1 mL of complete medium. Gently swirl the plate.
- Incubation & Analysis: Incubate at 37°C for 6-24h, then replace medium. Assess knockdown efficiency (qRT-PCR/Western) at 48-72h. Evaluate functional immunomodulatory changes in co-culture assays.
Quantitative Data Summary: Lipofection Table 2: Performance Metrics of Lipofection in MSCs
| Parameter | Typical Range | Notes |
|---|---|---|
| Transfection Efficiency | 30-70% | Optimized for siRNA/mRNA; lower for plasmid DNA in MSCs. |
| Cell Viability | 80-95% | Generally higher viability than electroporation. |
| Optimal Cargo | siRNA, mRNA, small plasmids | Ideal for knockdown/overexpression screens. |
| Protocol Complexity | Low | Simple, amenable to high-throughput formats. |
| Cost per Reaction | Moderate | Reagent cost is primary factor. |
Transposon Systems for Stable Engineering
Transposon systems (e.g., Sleeping Beauty (SB), PiggyBac (PB)) enable genomic integration and long-term transgene expression via a "cut-and-paste" or "cut-and-copy" mechanism facilitated by transposase enzyme.
Key Application: Generation of stable, clonal MSC lines constitutively or inducibly expressing immunomodulatory transgenes (e.g., soluble TNFα receptor, HO-1) for preclinical in vivo therapy studies.
Protocol: Generating Stable MSC Lines using PiggyBac Transposon
- Vector Design: Use a bi-cistronic PB donor plasmid carrying the immunomodulatory gene (e.g., IL-10) and a selectable marker (e.g., puromycin resistance) flanked by PB Terminal Inverted Repeats (TIRs). Use a separate plasmid or mRNA for the PB transposase.
- Co-transfection: Co-transfect 1 x 10^6 MSCs via nucleofection (see Protocol 1) with a 1:3 mass ratio of transposase:donor plasmid (e.g., 1 µg transposase mRNA + 3 µg donor plasmid).
- Selection & Expansion: 48 hours post-transfection, begin selection with puromycin (0.5-2 µg/mL, dose determined by kill curve). Maintain selection for 7-14 days until distinct colonies form.
- Clonal Isolation & Validation: Pick colonies using cloning rings. Expand clonal lines and validate transgene integration (genomic PCR, Southern blot), expression (flow cytometry, ELISA), and sustained immunomodulatory function (e.g., T-cell proliferation assay).
Quantitative Data Summary: Transposon Systems Table 3: Performance Metrics of Transposon Systems in MSCs
| Parameter | Sleeping Beauty | PiggyBac | Notes |
|---|---|---|---|
| Integration Mechanism | Cut-and-paste | Cut-and-copy | PB allows re-mobilization. |
| Integration Site Preference | TA dinucleotide (random) | TTAA tetranucleotide (random) | Both relatively random vs. lentivirus. |
| Cargo Capacity | High (>10 kbp) | Very High (>100 kbp) | PB superior for large constructs. |
| Stable Colony Formation | 2-4 weeks | 2-4 weeks | Requires antibiotic selection. |
| Long-Term Expression | Months in vitro & in vivo | Months in vitro & in vivo | Enables persistent immunomodulation. |
Diagram 2: Stable MSC line generation using PiggyBac transposon.
The Scientist's Toolkit: Key Reagent Solutions
Table 4: Essential Materials for Non-Viral MSC Transfection
| Reagent/Material | Function/Description | Example Product |
|---|---|---|
| Human MSC Nucleofector Kit | Cell-type specific buffer/electroporator programs for high-efficiency MSC transfection. | Lonza, VPE-1001 |
| Lipofectamine 3000 or RNAiMAX | Cationic lipid reagents optimized for plasmid DNA or siRNA/mRNA delivery, respectively. | Thermo Fisher Scientific |
| PiggyBac Transposon System | Donor vectors and transposase for high-cargo capacity, stable genomic integration. | System Biosciences (PB210PA-1) |
| Endotoxin-Free Plasmid Prep Kits | Ensures high-purity DNA to minimize cytotoxicity and immune activation in MSCs. | Qiagen EndoFree Plasmid Kits |
| Puromycin Dihydrochloride | Selection antibiotic for isolating stably transduced MSC clones post-transfection. | Thermo Fisher Scientific |
| Opt-MEM I Reduced Serum Medium | Low-serum medium used for diluting lipids/nucleic acids during lipoplex formation. | Thermo Fisher Scientific |
| MSC-Functional Media | Specialized, serum-free media supporting MSC health post-transfection for functional assays. | STEMCELL Technologies, MesenCult-ACF |
Within the thesis on genetic modification of mesenchymal stromal cells (MSCs) to enhance immunomodulation, precise gene editing is paramount. Knock-out (KO) strategies aim to disrupt genes encoding negative regulators (e.g., PD-L1, SOCS1) to potentiate MSC immunosuppressive function. Conversely, knock-in (KI) strategies enable targeted insertion of therapeutic transgenes (e.g., IL-10, HO-1, IFN-γ receptors) into safe-harbor loci (e.g., AAVS1), ensuring stable and enhanced immunomodulatory protein expression. CRISPR-Cas9 remains the foundational tool, but emerging editors like base and prime editors offer refined precision for single nucleotide changes without double-strand breaks (DSBs), crucial for modeling or correcting specific single nucleotide polymorphisms (SNPs) relevant to MSC potency.
Table 1: Comparison of Key Gene Editing Platforms for MSC Immunomodulation Engineering
| Platform | Editing Type | DSB Required? | Primary Use in MSC Context | Typical Efficiency in MSCs* | Key Limitation |
|---|---|---|---|---|---|
| CRISPR-Cas9 Nuclease | KO, KI via HDR | Yes | Disruption of immunoinhibitory genes (e.g., IFNγR1 KO). | KO: 20-80% (varies); KI: 1-20% | Off-target effects; low HDR efficiency. |
| CRISPR-Cas9 Nickase (D10A) | KI, precise edits | No (nicks) | Safer insertion of immunomodulatory cassettes. | KI: 5-30% | Requires paired gRNAs; lower efficiency than nuclease. |
| Base Editor (BE4) | C•G to T•A, A•T to G•C | No | Introducing or correcting SNPs in immunomodulatory genes (e.g., IL-10 promoter). | 10-50% (base change) | Target sequence context limitations; bystander edits. |
| Prime Editor (PE2) | All 12 base substitutions, small insertions/deletions | No | Precise installation of point mutations in cytokine receptors. | 1-30% | Lower efficiency; complex gRNA design. |
| CRISPR-Cas9 with Recruiting Peptides (e.g., p65, HSF1) | Targeted transcriptional activation (CRISPRa) | No | Overexpression of endogenous immunomodulatory genes (e.g., IDO1, PTGS2). | 5-50x mRNA upregulation | Transient effect; requires fusion protein. |
*Efficiencies are highly dependent on MSC source, delivery method, and culture conditions.
Detailed Experimental Protocols
Protocol 1: Knock-out of PD-L1 in Human Bone Marrow-Derived MSCs using CRISPR-Cas9 RNP
Objective: Disrupt the CD274 (PD-L1) gene to enhance MSC immunomodulatory function in allogeneic settings. Materials: See "Scientist's Toolkit" below. Procedure:
- gRNA Design & Preparation: Design two gRNAs targeting early exons of human CD274. Synthesize crRNA and tracrRNA. Resuspend to 100 µM in nuclease-free duplex buffer. Mix equimolar crRNA and tracrRNA (e.g., 5 µL each), heat to 95°C for 5 min, and cool slowly to form gRNA duplex.
- RNP Complex Formation: For one nucleofection, mix 3 µg (≈ 18 pmol) of purified Cas9 protein with 1.5 µL of 100 µM gRNA duplex (150 pmol). Incubate at room temperature for 10 min.
- MSC Preparation: Culture primary human BM-MSCs to 80-90% confluence (P3-P5). Harvest using TrypLE. Count and pellet 2.0 x 10^5 cells.
- Nucleofection: Use the Lonza 4D-Nucleofector. Resuspend cell pellet in 100 µL of P3 Primary Cell Solution. Add the RNP complex. Transfer to a certified cuvette. Run program DS-150. Immediately add 500 µL of pre-warmed, antibiotic-free culture medium.
- Recovery & Expansion: Plate cells in a T-75 flask. Change medium after 24h. Expand for 3-5 days.
- Analysis: Harvest cells. Assess KO efficiency via flow cytometry (anti-PD-L1 antibody) and T7 Endonuclease I assay on genomic DNA. Clone by single-cell sorting and validate by Sanger sequencing.
Protocol 2: HDR-Mediated Knock-in of an IL-10 Expression Cassette into theAAVS1Safe Harbor Locus
Objective: Generate MSCs with stable, site-specific expression of human IL-10. Materials: See toolkit. Donor template is a plasmid or ssDNA containing a PGK-IL10-T2A-GFP-pA cassette flanked by 800 bp homology arms to the human AAVS1 locus. Procedure:
- gRNA Design & RNP Prep: Use a validated AAVS1-targeting gRNA. Form RNP as in Protocol 1, step 2.
- Donor Template Delivery: Include 2 µg of supercoiled donor plasmid or 200 pmol of ssDNA donor with the RNP complex during nucleofection.
- Nucleofection & Recovery: Follow Protocol 1, steps 3-5, using 1.0 x 10^5 cells.
- Enrichment & Cloning: After 48h, begin puromycin selection (0.5 µg/mL) for 7-10 days. FACS sort GFP+ cells for clonal expansion.
- Genomic Validation: Perform junction PCR (one primer in genomic locus outside homology arm, one in the insert) and Sanger sequencing. Confirm IL-10 secretion via ELISA.
Visualizations
Diagram 1: Core CRISPR Strategies for MSC Engineering
Diagram 2: MSC Gene Editing Workflow
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Chemically Modified sgRNA (synthetized) | Enhanced nuclease stability and RNP formation efficiency compared to in vitro transcribed guides. |
| Recombinant S. pyogenes Cas9 Nuclease (NLS-tagged) | High-purity, ready-to-use protein for RNP formation, reducing plasmid toxicity and off-target duration. |
| AAVS1 Safe Harbor Targeting Kit | Validated gRNA and donor template controls for reliable knock-in, ensuring consistent experimental baselines. |
| Lonza P3 Primary Cell 4D-Nucleofector Kit | Optimized buffer/nucleofector program for high efficiency (up to 80% KO) and viability in human MSCs. |
| T7 Endonuclease I | Fast, cost-effective assay for initial screening of indel efficiency at target genomic loci. |
| Recombinant Human SCF, FGF-2 | Essential for maintaining MSC stemness and proliferative capacity during post-editing recovery and expansion. |
| Puromycin Dihydrochloride | Selection antibiotic for cells with integrated resistance markers (e.g., from knock-in donor plasmids). |
| CloneR Supplement | Enhances single-cell survival and clonal outgrowth post-FACS sorting, critical for isolating isogenic lines. |
| HiFi Cas9 Protein | Engineered variant with significantly reduced off-target effects, crucial for clinically relevant edits. |
| PCR-based HDR Donor Template | Linear double-stranded DNA with homology arms; reduces random integration risk vs. plasmid donors. |
Within the thesis on "Genetic modification of MSCs to enhance immunomodulation," this document details application notes and protocols for engineering mesenchymal stromal cells (MSCs) to overexpress four prime therapeutic targets: Heme Oxygenase-1 (HO-1), Programmed Death-Ligand 1 (PD-L1), Interleukin-4 (IL-4), and Tumor Necrosis Factor Alpha (TNFα) Receptors. These modifications aim to significantly amplify the innate immunosuppressive, anti-inflammatory, and tissue-protective functions of MSCs for treating autoimmune diseases, graft-versus-host disease (GvHD), and inflammatory disorders.
Target Rationale & Application Notes
1. Heme Oxygenase-1 (HO-1)
- Role: Catalyzes degradation of pro-oxidant heme into biliverdin (antioxidant), carbon monoxide (anti-inflammatory, vasodilatory), and free iron. Potently suppresses T-cell proliferation, dendritic cell maturation, and pro-inflammatory cytokine production.
- Therapeutic Application: Engineered HO-1-overexpressing MSCs show enhanced survival in oxidative microenvironments (e.g., infarcted tissue) and superior efficacy in models of ischemia-reperfusion injury, colitis, and solid organ transplant rejection.
2. Programmed Death-Ligand 1 (PD-L1)
- Role: Binds to PD-1 receptor on activated T cells, delivering an inhibitory signal that induces T-cell exhaustion, anergy, or apoptosis.
- Therapeutic Application: PD-L1-overexpressing MSCs exhibit potent, contact-dependent suppression of T-cell-driven pathologies. Particularly promising in models of GvHD, rheumatoid arthritis, and inflammatory bowel disease (IBD), where they can directly dampen pathogenic T-cell clones.
3. Interleukin-4 (IL-4)
- Role: Canonical Th2 cytokine that drives anti-inflammatory macrophage polarization to the M2 phenotype, promotes B-cell class switching, and inhibits Th1/Th17 responses.
- Therapeutic Application: IL-4-secreting MSCs act as a local " cytokine sink and factory," redirecting the immune milieu towards a regulatory state. Effective in allergy models, asthma, and as an adjuvant in tissue repair by promoting alternative macrophage activation.
4. Tumor Necrosis Factor Alpha (TNFα) Receptors (TNFR1/2)
- Role: Soluble or membrane-bound decoy receptors (e.g., sTNFR1, sTNFR2) bind and neutralize excess TNFα, a key driver of inflammation in diseases like rheumatoid arthritis and Crohn's disease.
- Therapeutic Application: MSCs engineered to overexpress TNFRs function as high-affinity, local TNFα antagonists. This protects the MSCs themselves from TNFα-induced apoptosis and provides targeted, sustained anti-cytokine therapy, surpassing the pharmacokinetic limitations of systemic biologic drugs.
Table 1: Summary of Key Findings from Preclinical Studies on Genetically Modified MSCs
| Target | Disease Model | Key Metric | Control MSC Result | Engineered MSC Result | Reference (Sample) |
|---|---|---|---|---|---|
| HO-1 | Myocardial Infarction | Infarct Size Reduction | ~18% reduction | ~45% reduction | Circ. Res. 2021 |
| PD-L1 | Acute GvHD | Mouse Survival (Day 60) | 40% survival | 90% survival | Blood 2022 |
| IL-4 | Collagen-Induced Arthritis | Clinical Arthritis Score | Score: 8.5 | Score: 3.2 | Stem Cells 2023 |
| TNFR2 | Dextran Sulfate Colitis | Disease Activity Index | Score: 9.1 | Score: 4.3 | Sci. Transl. Med. 2020 |
Detailed Experimental Protocols
Protocol 1: Lentiviral Vector Construction & MSC Transduction Objective: Stably overexpress HO-1, PD-L1, IL-4, or TNFR2 in human bone marrow-derived MSCs. Materials: See Scientist's Toolkit. Procedure:
- Gene Cloning: Subclone human HMOX1, CD274 (PD-L1), IL4, or TNFRSF1B (TNFR2) cDNA into a lentiviral transfer plasmid (e.g., pLVX-EF1α-IRES-Puro).
- Virus Production: Co-transfect HEK293T cells with the transfer plasmid, psPAX2 (packaging), and pMD2.G (VSV-G envelope) plasmids using polyethylenimine (PEI).
- Harvest & Concentration: Collect lentivirus-containing supernatant at 48h and 72h post-transfection. Concentrate via ultracentrifugation (80,000 x g, 2h).
- MSC Transduction: Seed passage 3 MSCs at 10,000 cells/cm². Add viral supernatant with 8 µg/mL polybrene. Spinfect at 800 x g for 45 min at 32°C.
- Selection & Expansion: Begin puromycin selection (1–2 µg/mL) 72h post-transduction. Maintain for 1 week, then expand positive pools.
Protocol 2: In Vitro T-Cell Suppression Assay Objective: Validate the immunomodulatory function of engineered MSCs. Procedure:
- Prepare Effectors: Isolate human PBMCs from buffy coats. Label CD3⁺ T cells with CellTrace Violet (CTV).
- Activate T Cells: Stimulate CTV-labeled T cells with anti-CD3/CD28 beads (bead:cell ratio 1:1) in RPMI+10% FBS.
- Co-culture: Seed engineered or control MSCs in a 96-well plate (5,000 cells/well). After adherence, add activated T cells at MSC:T cell ratios of 1:5, 1:10, and 1:20.
- Analysis: After 72-96h, harvest cells and analyze by flow cytometry.
- Proliferation: Measure CTV dilution in CD3⁺ gate.
- Activation: Stain for CD25 and CD69.
- Apoptosis: Stain with Annexin V/PI.
Signaling Pathways & Workflows
Diagram Title: HO-1 Mediated Immunomodulatory Mechanism
Diagram Title: Engineered MSC Development Workflow
The Scientist's Toolkit
Table 2: Essential Research Reagents & Materials
| Item | Function/Application | Example Vendor/Cat. No. |
|---|---|---|
| Lentiviral Packaging Mix (psPAX2/pMD2.G) | Essential plasmids for producing 2nd/3rd generation lentiviral particles. | Addgene #12260, #12259 |
| Polyethylenimine (PEI), Linear | High-efficiency transfection reagent for 293T cells during virus production. | Polysciences #23966 |
| Polybrene (Hexadimethrine bromide) | Cationic polymer that enhances viral transduction efficiency by neutralizing charge repulsion. | Sigma-Aldrich H9268 |
| Puromycin Dihydrochloride | Selection antibiotic for cells transduced with puromycin resistance-containing vectors. | Thermo Fisher A1113803 |
| Recombinant Human TNF-α | Critical for in vitro challenge assays to test TNFR-modified MSC function and survival. | PeproTech #300-01A |
| Anti-human CD274 (PD-L1) APC Antibody | Flow cytometry validation of surface PD-L1 overexpression on engineered MSCs. | BioLegend #329708 |
| CellTrace Violet Cell Proliferation Kit | Fluorescent dye to track and quantify T-cell proliferation in suppression assays. | Thermo Fisher C34557 |
| Human IL-4 ELISA Kit | Quantification of IL-4 secretion from engineered MSCs in supernatant. | R&D Systems D4050 |
| Human HO-1/HMOX1 ELISA Kit | Quantification of intracellular and secreted HO-1 protein levels. | Enzo ADI-960-071 |
Navigating the Lab: Solving Critical Challenges in Safety, Efficacy, and Scale-Up
Application Notes
Genetic modification of Mesenchymal Stromal Cells (MSCs) to enhance their immunomodulatory potential is a cornerstone of advanced cell therapy research for conditions like graft-versus-host disease (GvHD) and autoimmune disorders. The primary tools for stable genetic modification involve integrating vectors, such as lentiviruses and transposon systems. However, the associated risks of insertional mutagenesis (oncogene activation or tumor suppressor disruption due to random vector integration) and off-target effects from genome-editing nucleases like CRISPR-Cas9 pose significant safety hurdles for clinical translation. This document outlines current strategies, quantitative risk assessments, and detailed protocols to mitigate these risks, framed within the context of MSC immunomodulation engineering.
Quantitative Data on Integration and Editing Risks
Table 1: Comparison of Genetic Modification Tools for MSCs: Risks and Mitigation
| Tool/System | Primary Risk | Reported Unwanted Event Frequency (Range) | Key Mitigation Strategy |
|---|---|---|---|
| Lentiviral Vector | Insertional Mutagenesis | Clonal dominance in preclinical models: 5-15% (varies with design, titer). | Use of Self-Inactivating (SIN) designs, chromatin insulators (cHS4). |
| Sleeping Beauty Transposon | Insertional Mutagenesis | Lower genotoxicity profile vs. LV; exact frequency context-dependent. | Use of hyperactive, high-fidelity transposase (SB100X); codon optimization. |
| CRISPR-Cas9 (Knock-in) | Off-target editing & On-target mutations | Off-target indels: Varies widely (0.1% to >50%) based on guide design. | High-fidelity Cas9 variants (e.g., SpCas9-HF1); careful gRNA design & validation. |
| CRISPR Base/Prime Editors | Off-target & bystander edits | Off-target editing generally <0.1%; bystander edits can be higher. | Use of latest editor versions (PE3, PE5); optimized pegRNA design. |
Table 2: Key Assays for Risk Assessment in Modified MSCs
| Assay | Purpose | Readout | Acceptable Threshold (Example) |
|---|---|---|---|
| Linear Amplification-Mediated (LAM)-PCR / NGS | Map vector integration sites genome-wide. | Identification of genomic loci near oncogenes (e.g., LM02). | No integrations within 50kb of known oncogenes in final product clone. |
| GUIDE-seq / CIRCLE-seq | Identify CRISPR-Cas9 off-target sites genome-wide. | List of potential off-target loci with indel frequencies. | No off-target indels detected above 0.1% frequency at in silico-predicted sites. |
| Targeted Deep Sequencing | Quantify on-target editing efficiency & purity. | Percentage of intended HDR vs. indel (NHEJ) alleles. | >80% HDR for knock-in; <5% NHEJ at on-target site. |
| Long-term Clonogenic Assay | Assess proliferative dominance indicative of transformation risk. | Growth kinetics and morphology of single-cell-derived clones. | No evidence of hyper-proliferative or morphologically aberrant clones. |
Experimental Protocols
Protocol 1: Integration Site Analysis (ISA) via LAM-PCR and NGS for Lentiviral-Modified MSCs Objective: To map and analyze the genomic distribution of lentiviral vector integration sites in a polyclonal or clonal population of engineered MSCs. Materials: DNA from modified MSCs, LV-specific biotinylated primer, linker cassette, T4 DNA ligase, magnetic streptavidin beads, PCR reagents, NGS library prep kit. Procedure:
- Digestion & Ligation: Digest 1µg of genomic DNA with a frequent-cutter restriction enzyme (e.g., MseI). Ligate a known double-stranded linker cassette to the digested ends.
- Linear Amplification: Perform a linear PCR using a biotinylated primer specific to the LTR region of the integrated vector.
- Capture & 2nd Strand Synthesis: Capture the biotinylated single-stranded DNA product on streptavidin magnetic beads. Synthesize the second strand on-bead.
- Exponential PCR: Elute the double-stranded DNA and perform a nested PCR using a vector-specific primer and a linker-specific primer.
- NGS Library Preparation & Analysis: Purify the PCR product, prepare an NGS library, and sequence. Map sequencing reads to the human reference genome (hg38) using specialized software (e.g., LVcaller, INSPIIRED). Analyze for proximity to transcription start sites, oncogenes, and genomic safe harbors.
Protocol 2: Off-Target Assessment for CRISPR-Cas9 Edited MSCs using GUIDE-seq Objective: To empirically determine genome-wide off-target sites of a given CRISPR-Cas9 ribonucleoprotein (RNP) complex in MSCs. Materials: MSC cells, Cas9 protein, target-specific sgRNA, GUIDE-seq oligonucleotide (dsODN), transfection reagent (e.g., nucleofection kit), genomic DNA extraction kit, PCR reagents, NGS platform. Procedure:
- Co-transfection: Co-deliver the Cas9 RNP complex and the dsODN (a short, blunt, double-stranded oligo tag) into MSCs via nucleofection.
- Genomic DNA Extraction: Culture cells for 72 hours, then extract high-molecular-weight genomic DNA.
- Library Preparation for Sequencing: Perform two nested PCRs. The first uses a primer specific to the dsODN tag and a primer for a known on-target locus (positive control). The second, nested PCR adds Illumina adapters and sample indices.
- Sequencing & Bioinformatic Analysis: Sequence the libraries on a high-throughput platform. Use the GUIDE-seq computational pipeline to align reads, identify dsODN integration sites, and call off-target loci. Validate top candidate sites by targeted amplicon sequencing.
Protocol 3: Clonal Analysis for Insertional Mutagenesis Risk Objective: To screen single-cell-derived clones of genetically modified MSCs for aberrant growth indicative of insertional activation. Materials: Modified MSC pool, 96-well plates, conditioned media, cell staining solution, genomic DNA extraction kit. Procedure:
- Single-Cell Sorting: Using FACS, sort single modified MSCs into individual wells of a 96-well plate containing conditioned medium.
- Clonal Expansion: Expand clones over 4-6 weeks, monitoring morphology and growth rate.
- Phenotypic Screening: Identify clones with significantly accelerated proliferation or altered morphology (e.g., loss of contact inhibition).
- Molecular Analysis: Perform ISA (Protocol 1) on both normally growing and hyper-proliferative clones to correlate growth phenotype with integration site.
The Scientist's Toolkit
Table 3: Essential Research Reagents for Risk Mitigation Studies
| Item | Function in Risk Mitigation | Example Product/Type |
|---|---|---|
| High-Fidelity Cas9 Variant | Reduces off-target editing while maintaining on-target activity. | SpCas9-HF1, eSpCas9(1.1) |
| Self-Inactivating (SIN) Lentiviral Vector | Eliminates viral promoter activity in integrated provirus, lowering genotoxicity. | 3rd/4th Generation LV systems with deleted U3 region. |
| Chromatin Insulator | Can shield surrounding genes from enhancer effects of the integrated vector. | cHS4 core insulator element. |
| Hyperactive Transposase | Increases integration efficiency, allowing lower DNA amounts, potentially favoring safer integrations. | Sleeping Beauty SB100X transposase. |
| Genomic Safe Harbor Site gRNAs | Targets integration or knock-in to loci with minimal disruption risk (e.g., AAVS1, CCR5). | Validated sgRNAs for human AAVS1 (PPP1R12C) locus. |
| All-in-One NGS Off-target Kit | Streamlines detection of CRISPR off-targets and integration sites. | Commercial GUIDE-seq or CIRCLE-seq kits. |
| Recombinant Cas9 Nuclease | For RNP delivery, which reduces off-target risk vs. plasmid DNA and allows precise dosing. | Purified S. pyogenes Cas9 protein. |
Visualizations
Title: Workflow for Integration Site Analysis (LAM-PCR-NGS)
Title: Off-target Detection via GUIDE-seq Workflow
Title: Risk Mitigation Strategy Logic for Engineered MSCs
Within the broader thesis on the genetic modification of mesenchymal stromal cells (MSCs) to enhance immunomodulation, precise control over transgene expression is paramount. Inducible promoters and genetic safety switches enable temporal, dose-dependent control of therapeutic transgenes (e.g., anti-inflammatory cytokines) and provide a fail-safe mechanism to eliminate engineered cells if needed, enhancing both experimental rigor and clinical safety.
Application Notes
Tetracycline-Inducible Systems for Cytokine Expression
The Tet-On system allows doxycycline-dependent expression of immunomodulatory factors like IL-10 or TGF-β in engineered MSCs. This enables researchers to correlate transgene dosage with functional outcomes in co-culture assays with immune cells.
Rapamycin-Induced Dimerization Switches
Chemically-induced dimerization systems offer rapid, reversible control of signaling pathways. This is particularly useful for activating synthetic receptors (e.g., engineered Notch) in MSCs to secrete therapeutic payloads in response to a small molecule.
Safety Switches for Clinical Translation
The inclusion of a suicide gene, such as inducible caspase 9 (iCasp9), activated by a clinical-grade small molecule (AP1903/Rimiducid), is a critical safety contingency for adoptive cell therapies. This allows for the ablation of engineered MSCs in case of adverse events.
Experimental Protocols
Protocol 1: Validating a Tetracycline-Inducible Promoter in MSCs
Objective: To assess the dynamic range and leakage of a Tet-On 3G system driving an IL-10 transgene in human bone marrow-derived MSCs.
Materials:
- Lentiviral vectors pLVX-Tet3G (transactivator) and pLVX-TRE3G-IL10-Puro (response).
- Human BM-MSCs (passage 3-5).
- Doxycycline hyclate stock (1 mg/mL in PBS).
- Puromycin dihydrochloride.
- IL-10 ELISA kit.
Procedure:
- Viral Transduction: Day 1: Seed MSCs at 10,000 cells/cm². Day 2: Transduce with pLVX-Tet3G virus at MOI 5 in the presence of 8 µg/mL polybrene. Spinfect at 800 × g for 45 min at 32°C.
- Selection: 48 hours post-transduction, begin selection with 1 µg/mL puromycin for 7 days to generate stable Tet3G-MSCs.
- Second Transduction: Transduce Tet3G-MSCs with pLVX-TRE3G-IL10-Puro virus (MOI 10). Select with a higher puromycin dose (2 µg/mL) for 5 days.
- Induction & Quantification: Seed dual-transduced MSCs in 24-well plates (20,000 cells/well). Treat with a doxycycline dose range (0, 10, 100, 1000 ng/mL) for 48 hours. Collect supernatant and quantify IL-10 via ELISA. Perform qPCR on cell lysates for IL-10 mRNA.
- Data Analysis: Calculate fold-induction (ON/OFF) and EC50 for doxycycline.
Protocol 2: Testing an iCasp9 Safety SwitchIn Vitro
Objective: To demonstrate efficient elimination of engineered MSCs upon addition of the dimerizing drug AP1903.
Materials:
- MSCs transduced with lentivector expressing iCasp9 and a reporter (e.g., GFP).
- AP1903 (Rimiducid) stock solution (10 µM in DMSO).
- Annexin V/7-AAD apoptosis detection kit.
- Real-time cell analyzer (e.g., xCELLigence) or bright-field microscope.
Procedure:
- Cell Seeding: Seed iCasp9-MSCs in triplicate in a 96-well plate (5,000 cells/well). Allow to adhere overnight.
- Drug Addition: Add AP1903 to final concentrations of 0, 1, 10, and 100 nM.
- Viability Monitoring:
- Method A (Real-time): Place plate on xCELLigence system to monitor cell index every 15 minutes for 72 hours.
- Method B (Endpoint): At 24 and 48 hours post-AP1903, stain cells with Annexin V/7-AAD and analyze by flow cytometry.
- Analysis: Determine the percentage of apoptotic/necrotic cells (Annexin V+/7-AAD+) and the concentration of AP1903 required for >95% elimination within 24 hours.
Data Presentation
Table 1: Performance Metrics of Common Inducible Systems for MSCs
| System | Inducer | Typical Induction Fold-Change | Time to Max Expression | Basal Leakage | Primary Use |
|---|---|---|---|---|---|
| Tet-On 3G | Doxycycline | 100 - 1,000x | 24 - 48 h | Very Low | Controlled cytokine expression |
| Cumate | Cumate | 50 - 200x | 24 - 48 h | Low | Gene function studies |
| iCasp9 | AP1903 | N/A (Cell Death) | Apoptosis in 2-8 h | None | Safety/ablation |
| RU486 (GeneSwitch) | Mifepristone | 10 - 50x | 12 - 24 h | Moderate | In vivo applications |
Table 2: Example IL-10 Induction Data from Tet-On MSCs (n=3)
| Doxycycline (ng/mL) | IL-10 Secretion (pg/mL/24h) | SD | Fold Induction vs. 0 ng/mL |
|---|---|---|---|
| 0 (OFF state) | 15.2 | ± 3.1 | 1.0 |
| 10 | 520.5 | ± 45.2 | 34.2 |
| 100 | 4,850.7 | ± 210.8 | 319.1 |
| 1000 | 12,150.3 | ± 505.5 | 799.4 |
Visualizations
Diagram 1: Tetracycline (Tet-On) Inducible System Mechanism
Diagram 2: iCasp9 Safety Switch Activation Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Inducible Expression in MSC Research
| Reagent/Category | Example Product | Function in Experiment |
|---|---|---|
| Inducible System Vectors | pLVX-Tet3G & pLVX-TRE3G (Takara Bio), pAC94-iCasp9-P2A-GFP (Addgene) | Core genetic components for building inducible or safety switch constructs. |
| Small Molecule Inducers | Doxycycline hyclate (Sigma D9891), AP1903/Rimiducid (MedChemExpress) | Pharmacologic triggers to turn on transgene expression or induce apoptosis. |
| Lentiviral Packaging Mix | Lenti-X Packaging Single Shots (Takara Bio) | For safe, efficient production of VSV-G pseudotyped lentivirus to transduce MSCs. |
| MSC-Specific Media | StemMACS MSC Expansion Media (Miltenyi) | Maintains MSC phenotype and multipotency during genetic manipulation and expansion. |
| Apoptosis Detection Kit | Annexin V-FITC/7-AAD Kit (BioLegend) | Gold-standard for quantifying early/late apoptosis after safety switch activation. |
| Cytokine Quantification | Human IL-10 DuoSet ELISA (R&D Systems) | Precise measurement of induced immunomodulatory factor secretion. |
This application note details methodologies central to a thesis on the genetic modification of mesenchymal stromal cells (MSCs) to enhance their immunomodulatory potential for therapeutic applications. The primary challenge lies in optimizing the in vivo dose and persistence of these engineered cells to achieve sustained efficacy while ensuring eventual clearance to mitigate long-term risks. This document provides current protocols and data analysis frameworks for quantifying these critical parameters.
Key Quantitative Data: Engineered MSC Therapies
Table 1: Comparative In Vivo Persistence of MSC Variants
| MSC Type / Modification | Animal Model | Route of Administration | Initial Dose (cells) | Half-life (Days) | Peak Engraftment (% at site) | Key Measured Outcome |
|---|---|---|---|---|---|---|
| Naive (Unmodified) MSCs | NSG Mouse (IV) | Intravenous | 1 x 10^6 | 2-4 | < 0.1% (Lung) | Rapid clearance via lung sequestration |
| MSCs Expressing CXCR4 | NSG Mouse (IV) | Intravenous | 1 x 10^6 | 5-7 | 1.2% (Bone Marrow) | Enhanced homing to bone marrow niche |
| MSCs Overexpressing Bcl-2 | C57BL/6 Mouse (IV) | Intravenous | 1 x 10^6 | 10-14 | 0.8% (Spleen) | Reduced apoptosis, extended systemic presence |
| MSCs with HIF-1α KO (CRISPR) | NSG Mouse (IM) | Intramuscular | 5 x 10^5 | >30 | 15% (Muscle) | Enhanced survival in hypoxic tissue |
| MSC-derived EV (Extracellular Vesicles) | Rat, MI Model | Intra-myocardial | 1 x 10^10 particles | 0.2 | N/A | Rapid clearance, repeated dosing required |
Table 2: Correlation Between Dose, Persistence, and Immunomodulatory Efficacy in GvHD Model
| Dose (cells/mouse) | Serum IL-10 Peak (pg/mL) | Treg Induction (%) | Survival Rate (Day 60) | Bioluminescence Signal Duration (Days) |
|---|---|---|---|---|
| 2.5 x 10^5 | 45 ± 12 | 8.2 ± 1.1 | 40% | 7 ± 2 |
| 5 x 10^5 | 98 ± 22 | 12.5 ± 2.3 | 70% | 14 ± 3 |
| 1 x 10^6 | 210 ± 45 | 18.7 ± 3.1 | 90% | 21 ± 4 |
| 2 x 10^6 | 205 ± 40 | 19.1 ± 2.8 | 85% | 28 ± 5* |
Note: *Higher dose associated with microvasculature occlusion incidents.
Detailed Experimental Protocols
Protocol 1: Longitudinal Tracking of Genetically Modified MSC Persistence Using Bioluminescence Imaging (BLI)
Objective: To non-invasively quantify the in vivo biodistribution and persistence of luciferase-expressing MSCs.
Materials:
- Firefly luciferase (FLuc)-expressing MSCs (modified/unmodified).
- D-Luciferin, potassium salt (15 mg/mL in PBS).
- In vivo imaging system (IVIS) or equivalent.
- Anesthetic (isoflurane or ketamine/xylazine mix).
- Immunodeficient or syngeneic mice.
Procedure:
- Cell Preparation: Harvest engineered MSCs at passage 3-5. Confirm >95% viability via trypan blue exclusion.
- Administration: Inject a defined dose (e.g., 1x10^6 cells in 100µL PBS) via the intended route (IV, IP, IM) into animals (n=5-8 per group).
- Imaging Time Course: Image at 4 hours post-injection (initial engraftment), then daily for 7 days, followed by weekly until signal returns to background.
- Substrate Injection: Inject D-Luciferin intraperitoneally (150 mg/kg) 10 minutes prior to imaging.
- Image Acquisition: Anesthetize animals, place in imaging chamber. Acquire grayscale and luminescent images (exposure: 1-60 sec, binning: medium).
- Data Analysis: Use imaging software to draw regions of interest (ROIs) around the signal. Quantify total flux (photons/sec). Plot flux vs. time to calculate pharmacokinetic half-life.
Protocol 2: Ex Vivo qPCR-Based Quantification of Human-SpecificAluSequences for Engraftment
Objective: To precisely quantify human MSC engraftment in rodent tissues with high sensitivity, independent of reporter genes.
Materials:
- Tissue samples (spleen, lung, liver, bone marrow).
- DNA extraction kit (e.g., DNeasy Blood & Tissue).
- Primers for human Alu sequences (F: 5'-ACGCCTGTAATCCCAGCACTT-3'; R: 5'-TCGCCCAGGCTGGAGTGCA-3').
- Primers for mouse Gapdh as control.
- SYBR Green qPCR Master Mix.
- Real-time PCR system.
Procedure:
- Tissue Harvest: At predetermined endpoints, perfuse animals with PBS. Harvest and weigh target organs.
- Genomic DNA Extraction: Homogenize tissues, extract total DNA per kit instructions. Quantify DNA using a Nanodrop.
- Standard Curve Preparation: Extract DNA from a known number of human MSCs (e.g., 10 to 1,000,000 cells). Serially dilute and use to generate a standard curve of Alu Ct vs. log(cell number).
- qPCR Setup: Perform reactions in triplicate: 10µL SYBR mix, 0.5µM primers, 50ng sample DNA. Cycling: 95°C for 10 min; 40 cycles of 95°C/15s, 60°C/60s.
- Quantification: From the standard curve, interpolate the number of human cells per organ. Normalize to organ weight.
Protocol 3: Functional Assessment ofIn VivoImmunomodulation
Objective: To evaluate the functional efficacy of persistent, engineered MSCs.
Materials:
- Single-cell suspension from spleen/lymph nodes.
- Flow cytometry antibodies: CD4, CD25, Foxp3 (Tregs); CD4, IFN-γ (Th1); CD4, IL-17 (Th17).
- Intracellular cytokine staining kit.
- ELISA kits for TGF-β, IL-10, PGE2.
Procedure:
- Experimental Model: Utilize a disease model (e.g., GvHD, EAE) or immune challenge.
- Cell Harvest: 7-14 days post-MSC administration, harvest lymphoid organs.
- Immune Cell Profiling:
- For surface markers, stain single-cell suspensions, analyze via flow cytometry.
- For cytokines, stimulate cells with PMA/ionomycin + brefeldin A for 4-6h, perform intracellular staining.
- Soluble Mediator Analysis: Collect serum or plasma. Measure concentrations of immunomodulatory factors (IL-10, TGF-β) via ELISA per manufacturer protocol.
- Correlation: Correlate immune marker changes with persistence data from Protocol 1 or 2.
Visualizations
Title: PK/PD Pathway of Engineered MSC Therapies
Title: Workflow for Optimizing MSC Dose & Persistence
Title: Balance: Persistence Drives Efficacy vs. Risk
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Engineered MSC Persistence Studies
| Item / Reagent | Function in Research | Example Product/Catalog |
|---|---|---|
| Lentiviral Vector (Luc2/Puro) | Stable integration of bioluminescence reporter for in vivo tracking. | pCDH-EF1-Luc2-Puro (System Biosciences) |
| Human Alu Sequence qPCR Assay | Sensitive, species-specific quantification of human cell engraftment in tissues. | HsAluSq (QIAGEN, #PPH60050F) |
| D-Luciferin, Potassium Salt | Substrate for firefly luciferase, injected for BLI. | GoldBio, #LUCK-1G |
| Annexin V Apoptosis Kit | Assess ex vivo MSC apoptosis resistance post-modification (e.g., Bcl-2). | FITC Annexin V/Dead Cell Kit (Thermo, #V13242) |
| Recombinant Fibronectin/CH-296 | Coating for in vitro assays to mimic niche adhesion and study homing (CXCR4). | RetroNectin (Takara, #T100B) |
| Mouse/Rat Cytokine Multiplex ELISA | Profile broad panel of immunomodulatory cytokines from serum/tissue lysate. | LEGENDplex (BioLegend) |
| FOXP3 / Transcription Factor Staining Buffer Set | For intracellular staining of Tregs and other key immune cell populations. | Thermo Fisher, #00-5523-00 |
| G-CSF & SDF-1α ELISA Kits | Quantify niche factors influencing MSC homing and retention. | DuoSet ELISA (R&D Systems) |
| CRISPR-Cas9 Kit (HIF-1α) | Knockout genes to enhance survival in hypoxic target tissues. | Edit-R CRISPR-Cas9 (Horizon Discovery) |
| Cell Trace Violet (CTV) | Label MSCs for ex vivo flow cytometric tracking of division and persistence. | Thermo Fisher, #C34557 |
The transition from research-grade to Good Manufacturing Practice (GMP)-compliant clinical-scale production of genetically engineered Mesenchymal Stromal Cells (MSCs) presents significant, multifaceted hurdles. Within a thesis on enhancing MSC immunomodulation via genetic modification, these hurdles directly impact the viability, potency, and safety of the final cell therapy product. The core challenges span vector production, cell processing, quality control, and regulatory compliance.
Table 1: Key Hurdles in Scaling Engineered MSC Production
| Development Phase | Research-Grade Practice | GMP Clinical-Scale Requirement | Primary Challenge |
|---|---|---|---|
| Vector Production | Use of research-grade plasmids & packaging cells; transient transfection. | GMP-grade plasmids, master cell banks; validated, scalable production (e.g., stable producer lines). | Elimination of adventitious agents; consistent high-titer, pure vector batches. |
| Genetic Modification | Low-efficiency methods (e.g., electroporation); use of research-grade reagents. | Closed, automated systems (e.g., electroporation); defined, xeno-free reagents. | Achieving high viability & transduction efficiency at scale; process validation. |
| Cell Expansion | Open culture (flasks), FBS-containing media, manual passaging. | Closed bioreactors (e.g., hollow-fiber, stirred-tank), xeno-free media, automated feeding. | Maintaining critical quality attributes (phenotype, potency, karyotype) during rapid scale-up. |
| Quality Control (QC) | Limited in-process testing; research-only assays. | Extensive, validated release assays: potency, identity, purity, sterility, vector safety. | Developing quantitative, robust potency assays for immunomodulation; mycoplasma, endotoxin testing. |
| Characterization | Surface markers (minimal panel), proof-of-concept functional data. | Extended ISCT phenotype, genomic stability (karyotyping, SNP), biodistribution, tumorigenicity. | Demonstrating stable transgene expression without insertional mutagenesis or clonal dominance. |
Application Notes & Detailed Protocols
Protocol: GMP-Compliant Lentiviral Transduction of MSCs in a Closed System
Objective: To genetically modify human bone marrow-derived MSCs with a lentiviral vector encoding an immunomodulatory transgene (e.g., IL-10 or PD-L1) under GMP-like conditions suitable for scale-up.
Materials & Reagents (The Scientist's Toolkit):
Table 2: Key Research Reagent Solutions for GMP-Compliant Transduction
| Reagent/Material | Function | GMP-Compliant Sourcing Consideration |
|---|---|---|
| GMP-Grade MSC Medium | Xeno-free, serum-free cell expansion medium. | Eliminates batch variability and zoonotic risk from FBS. |
| Clinical-Grade LV Vector | Encoding therapeutic transgene and a reporter (e.g., GFP). | Produced under GMP using a Master Virus Bank; titered and tested for replication-competent lentivirus (RCL). |
| Polycationic Transduction Enhancer (e.g., protamine sulfate) | Increases viral adhesion to cell surface, improving efficiency. | Must be USP/EP grade for injection. |
| Closed System Cell Culture Bag or Bioreactor | Provides a sterile, scalable environment for cell growth and transduction. | Enables aseptic processing and reduces contamination risk (e.g., WAVE bioreactor, hollow-fiber system). |
| Automated Cell Processing System | For washing and concentration of cells post-transduction. | Maintains closed system (e.g., LOVO, Cytiva Sefia). |
| Validated QC Assay Reagents | For flow cytometry, ELISA, qPCR. | Assays must be qualified/validated for specificity, accuracy, precision. |
Detailed Methodology:
- MSC Thawing & Expansion:
- Thaw a validated GMP Master Cell Bank (MCB) vial of human MSCs in a 37°C water bath.
- Transfer cells to a pre-filled bag of warm GMP-grade MSC medium and centrifuge using a closed system cell processor.
- Seed cells at a density of 3,000 cells/cm² in a closed culture bag or bioreactor pre-equilibrated with medium. Maintain at 37°C, 5% CO2.
- Expand cells for 2-3 passages, monitoring population doublings and adherence to predefined phenotype (≥95% positive for CD73, CD90, CD105; ≤2% positive for CD45, CD34).
Pre-Transduction Preparation:
- Harvest cells at ~70% confluence using a GMP-grade, enzyme-free dissociation buffer in the closed system.
- Wash and resuspend cells in fresh, pre-warmed medium at a concentration of 1 x 10^6 cells/mL.
Transduction:
- In the closed culture bag, add clinical-grade protamine sulfate to the cell suspension at a final concentration of 5 µg/mL.
- Add the GMP-grade lentiviral vector at a pre-optimized Multiplicity of Infection (MOI) of 10-20. Gently mix.
- Seal the bag and place it on a rocking platform in the incubator (37°C, 5% CO2) for 16-24 hours.
Post-Transduction Processing:
- After incubation, transfer the bag to the closed cell processing system.
- Wash the cells twice with large volumes of PBS/medium to remove residual vector particles.
- Resuspend the cell pellet in fresh medium and return to a fresh culture bag/bioreactor for further expansion.
Analysis & Harvest:
- 48-72 hours post-transduction: Assess transduction efficiency via flow cytometry for the reporter gene (e.g., GFP). Expect 60-80% efficiency.
- Expand cells to the target final cell number (e.g., 1 x 10^9 cells for a clinical dose).
- Harvest using a closed-system detachment and washing protocol. Formulate in the final infusion solution (e.g., cryopreservation medium).
- Perform in-process and lot-release testing (see Table 3).
Protocol: Quantitative Potency Assay for IL-10-Engineered MSCs
Objective: To measure the immunosuppressive capacity of engineered MSCs via a standardized, quantitative co-culture assay simulating a pro-inflammatory environment.
Detailed Methodology:
- Peripheral Blood Mononuclear Cell (PBMC) Activation:
- Isolate PBMCs from healthy donor leukapheresis product using Ficoll density gradient centrifugation.
- Label PBMCs with a cell proliferation dye (e.g., CFSE).
- Activate PBMCs at 2 x 10^5 cells/well in a 96-well plate using a GMP-grade anti-CD3/CD28 activator (e.g., ImmunoCult) at a 1:50 dilution.
Co-Culture Setup:
- Seed irradiated (30 Gy) parental or IL-10-engineered MSCs in the same well at varying ratios (MSC:PBMC of 1:10, 1:20, 1:50). Use triplicates for each condition.
- Include controls: PBMCs alone (max proliferation), PBMCs + MSCs + activating agent (test), PBMCs + activating agent + high-dose immunosuppressant (e.g., cyclosporin A, inhibition control).
Incubation & Analysis:
- Incubate co-culture for 5 days at 37°C, 5% CO2.
- Harvest cells and stain for T-cell markers (CD3, CD4, CD8).
- Analyze by flow cytometry to measure:
- % Inhibition of Proliferation: Based on reduction of CFSE dilution in CD3+ T-cells.
- Cytokine Secretion: Collect supernatant and measure IFN-γ levels by ELISA (expected >70% reduction with potent engineered MSCs).
- IL-10 Output: Quantify human IL-10 in supernatant by ELISA to confirm transgene function.
Quality Control & Release Specifications
Table 3: Proposed Release Criteria for a Clinical Batch of Engineered MSCs
| Test Category | Specific Test | Acceptance Criterion | Method |
|---|---|---|---|
| Identity | Surface Marker Expression (ISCT) | ≥95% positive for CD73, CD90, CD105; ≤2% positive for CD45, CD34, CD19, CD11b, HLA-DR. | Flow Cytometry |
| Viability | Post-thaw Viability | ≥80% | Trypan Blue/Flow Cytometry (7-AAD) |
| Purity & Safety | Sterility (Bacteriology/Fungology) | No growth for 14 days. | BacT/ALERT |
| Mycoplasma | Negative. | PCR-based assay | |
| Endotoxin | <5.0 EU/kg/hr (or per dose limit). | LAL assay | |
| Replication-Competent Lentivirus (RCL) | Negative. | PCR/HeLa-based co-culture | |
| Potency | Transgene Expression (e.g., %GFP+) | ≥60% (or batch-specific range). | Flow Cytometry |
| Immunomodulation Assay | ≥50% inhibition of T-cell proliferation vs. control at 1:20 ratio. | PBMC Co-culture (see Protocol 2.2) | |
| Secreted Transgene Product (e.g., IL-10) | ≥1000 pg/10^6 cells/24h (batch-specific). | ELISA | |
| Genomic Safety | Copy Number Analysis | Consistent, defined range (e.g., 1-5 copies/cell). | ddPCR |
| Karyotype | Normal diploid complement (46, XY/XX). | G-Banding |
Visualizations
Title: GMP Workflow for Engineered MSC Manufacturing
Title: Engineered MSC Immunomodulation Pathways
Proof of Concept: Benchmarking Engineered MSCs Against Existing Therapies
This document provides application notes and protocols for advanced co-culture assays, developed within the broader thesis research on "Genetic Modification of MSCs to Enhance Immunomodulation." The central hypothesis posits that engineered MSCs overexpressing specific immunomodulatory factors (e.g., PD-L1, IL-10, TGF-β, HLA-G) exhibit superior suppression of immune cell activation and proliferation. These assays are critical for in vitro validation of these genetically modified (GM)-MSCs prior to in vivo or clinical evaluation. The protocols focus on quantifying the functional immunomodulatory capacity of GM-MSCs using peripheral blood mononuclear cells (PBMCs) and isolated immune cell subsets.
Table 1: Expected Immunomodulatory Effects of GM-MSCs in Co-culture Assays
| GM-MSC Modificaton | Target Immune Cell | Key Readout | Expected Inhibition vs. Wild-Type MSCs | Typical Assay Duration |
|---|---|---|---|---|
| PD-L1 Overexpression | Activated CD4+ T Cells | CFSE Proliferation (% divided) | 40-60% greater suppression | 5 days |
| IDO (Indoleamine 2,3-dioxygenase) Overexpression | PBMCs (Overall) | Tryptophan depletion (µM), Kynurenine production (µM) | 3-5 fold increase in kynurenine | 3 days |
| IL-10 Overexpression | Monocyte-derived DCs | Surface MHC-II Mean Fluorescence Intensity (MFI) | 50-70% reduction in MFI | 2 days |
| HLA-G1 Overexpression | NK Cells | CD107a Degranulation (% positive) | 60-80% greater inhibition | 18 hours |
| TGF-β1 Overexpression | CD4+ T Cells | Treg Induction (FoxP3+ % of CD4+) | 2-3 fold increase | 5 days |
Table 2: Essential Controls for Co-culture Validation
| Control Condition | Purpose | Acceptance Criterion for Valid Assay |
|---|---|---|
| Immune cells alone (no MSC) | Baseline activation/proliferation | Robust response to stimulus (e.g., >60% T cell proliferation) |
| Immune cells + Wild-Type MSCs | Baseline immunomodulation | Measurable suppression vs. "alone" control (e.g., 20-50% inhibition) |
| Immune cells + GM-MSCs | Test modified function | Statistically significant enhancement vs. Wild-Type MSCs (p<0.05) |
| GM-MSCs alone (no immune cells) | Assess factor secretion | Confirm transgene expression (ELISA/qPCR) |
Detailed Experimental Protocols
Protocol 3.1: Transwell Co-culture of GM-MSCs with Mitogen-Activated PBMCs
Objective: To assess the contact-independent immunosuppressive capacity of GM-MSCs via soluble factors. Materials: 24-well transwell plates (0.4 µm pore), GM-MSCs, fresh PBMCs from healthy donor, CFSE, anti-CD3/CD28 beads, flow cytometer.
- Seed GM-MSCs: Seed 5 x 10⁴ GM-MSCs or control MSCs in the lower chamber of the transwell plate in complete MSC medium. Allow to adhere overnight.
- Label PBMCs: Isolate PBMCs via density gradient centrifugation. Resuspend at 2 x 10⁶/mL in PBS/0.1% BSA. Add CFSE to a final concentration of 1 µM, incubate 10 min at 37°C. Quench with 5x volume of cold complete RPMI.
- Activate & Co-culture: Stimulate CFSE-labeled PBMCs (5 x 10⁵) with anti-CD3/CD28 beads (bead:cell ratio 1:1). Add the activated PBMCs to the upper transwell chamber.
- Harvest & Analyze: After 5 days, harvest PBMCs from the upper chamber. Analyze CFSE dilution in CD3+ T cells by flow cytometry. Use proliferation modeling software to calculate division index.
Protocol 3.2: Direct Co-culture with Isolated Immune Cell Subsets
Objective: To evaluate GM-MSC-mediated modulation of specific immune cell functions (e.g., NK cell cytotoxicity, DC maturation). Materials: GM-MSCs, isolated immune cell subsets (e.g., NK cells using CD56+ microbeads), maturation stimuli (LPS for DCs), flow cytometry antibodies. A. NK Cell Degranulation Assay:
- Seed MSCs: Seed 2 x 10⁴ GM-MSCs/well in a 96-well U-bottom plate.
- Isolate & Activate NK Cells: Isolate CD56+ NK cells from PBMCs. Pre-activate NK cells with IL-12 (10 ng/mL) and IL-15 (50 ng/mL) for 18 hours.
- Co-culture: Add pre-activated NK cells (2 x 10⁵) to the MSC monolayer (MSC:NK ratio 1:10). Add anti-CD107a antibody directly to the well. Incubate for 1 hour, then add monensin/Brefeldin A and incubate for an additional 5 hours.
- Analyze: Harvest cells, stain for CD56 and CD3, and analyze CD107a surface expression on CD3-CD56+ NK cells via flow cytometry.
Protocol 3.3: Cytokine Profiling & Metabolic Analysis
Objective: To quantify the secretome and metabolic changes induced by GM-MSC co-culture. Materials: Multiplex cytokine array (e.g., Luminex), HPLC/MS for metabolite analysis.
- Conditioned Media Collection: After 48-72 hours of co-culture (from Protocol 3.1 or 3.2), collect supernatants. Centrifuge to remove cells/debris.
- Analysis:
- Cytokines: Use a 25-plex human cytokine panel to quantify IL-2, IFN-γ, TNF-α, IL-10, IL-6, IL-4, etc.
- Metabolites: For IDO-overexpressing MSCs, analyze tryptophan and kynurenine levels via HPLC. Calculate the kynurenine/tryptophan ratio.
Visualized Workflows & Pathways
Diagram 1: GM-MSC Co-culture Experimental Workflow
Diagram 2: Key Immunomodulatory Pathways in GM-MSCs
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Advanced Co-culture Assays
| Reagent/Material | Supplier Examples | Function in Assay |
|---|---|---|
| Ficoll-Paque PLUS | Cytiva, Sigma-Aldrich | Density gradient medium for isolating PBMCs from whole blood. |
| CellTrace CFSE / Cell Proliferation Dyes | Thermo Fisher | Fluorescent cytoplasmic dye to track and quantify immune cell division by flow cytometry. |
| Human T-Activator CD3/CD28 Dynabeads | Thermo Fisher | Provides strong, uniform polyclonal stimulation of human T cells for activation control. |
| Human NK Cell Isolation Kit (CD56+) | Miltenyi Biotec, STEMCELL | Magnetic bead-based negative selection for high-purity untouched NK cell isolation. |
| LPS (Lipopolysaccharide) | Sigma-Aldrich, InvivoGen | Toll-like receptor agonist used to induce maturation in monocyte-derived dendritic cells. |
| Recombinant Human IL-2, IL-12, IL-15 | PeproTech, R&D Systems | Cytokines for pre-activation and maintenance of NK cells and T cell subsets. |
| Multiplex Cytokine Assay (e.g., LegendPlex) | BioLegend, Thermo Fisher | Bead-based immunoassay for simultaneous quantification of numerous cytokines from supernatant. |
| Anti-human CD107a (LAMP-1) FITC | BioLegend, BD Biosciences | Antibody for surface staining to measure degranulation of cytotoxic lymphocytes (NK cells, CD8+ T cells). |
| Transwell Permeable Supports (0.4 µm) | Corning, Falcon | Permits exchange of soluble factors while preventing cell contact between MSC and immune cell compartments. |
| FoxP3 / Transcription Factor Staining Buffer Set | Thermo Fisher | Permeabilization buffers for intracellular staining of key regulatory markers like FoxP3 (Tregs). |
Application Notes: Genetically Modified MSC Efficacy Across Immune-Mediated Disease Models
Within the thesis framework of enhancing MSC immunomodulation via genetic modification, selecting the appropriate preclinical model is paramount for validating therapeutic efficacy and mechanism. The following notes and data compare the performance of engineered MSCs (e.g., overexpressing IL-10, TGF-β, PD-L1, or HO-1) in four key immune-disease arenas.
Table 1: Efficacy Metrics of GM-MSCs in Preclinical Models
| Disease Model | Common Animal System | Key Efficacy Readouts (Quantitative) | Typical Efficacy Range of GM-MSCs vs. Wild-type MSCs |
|---|---|---|---|
| Graft-versus-Host Disease (GvHD) | MHC-mismatched murine allogeneic HCT (e.g., B6→B6D2F1) | Survival (%), Clinical GvHD Score, Histopathology (Grade), Serum Inflammatory Cytokines (pg/mL) | Survival: 70-90% vs. 40-60% (WT-MSC). IL-6/TNF-α reduction: 60-80% greater. |
| Rheumatoid Arthritis (RA) | Murine CIA (Collagen-Induced Arthritis) | Arthritis Score, Paw Swelling (mm), Histological Score, Bone Erosion (µCT), Anti-CII IgG (µg/mL) | Arthritis Score Reduction: 50-70% vs. 20-40%. Bone erosion: 40-60% improved protection. |
| Multiple Sclerosis (MS) | Murine EAE (Experimental Autoimmune Encephalomyelitis) | Clinical EAE Score, CNS Inflammatory Infiltrates (cells/mm²), Demyelination (%, Luxol Fast Blue), | Peak EAE Score Reduction: ~30-50% vs. ~10-25%. Cellular Infiltrate: 40-60% lower. |
| Inflammatory Bowel Disease (IBD) | Murine DSS-Induced Colitis or TNBS-Colitis | Disease Activity Index, Colon Length (cm), Histology Score, MPO Activity (U/g tissue) | DAI Improvement: 60-80% vs. 30-50%. Colon Length: 15-25% less shortening vs. control. |
Core Mechanistic Insights: GM-MSCs consistently show enhanced suppression of pathogenic T cells (Th1, Th17) and promotion of regulatory T cells (Tregs) across models, coupled with increased homing to inflamed sites and sustained paracrine factor secretion.
Detailed Experimental Protocols
Protocol 1: Efficacy Testing in a Murine Acute GvHD Model Objective: To assess the therapeutic potential of HO-1-overexpressing human MSCs (HO1-MSCs) in modulating lethal acute GvHD.
- HCT & GvHD Induction: Irradiate (e.g., 8.5 Gy) recipient B6D2F1 mice. Within 24h, infuse via tail vein with 5x10^6 bone marrow cells and 2x10^6 splenocytes from donor C57BL/6 mice.
- Therapeutic MSC Administration: On day +1 and +4 post-HCT, administer 1x10^6 HO1-MSCs or control MSCs in PBS via tail vein (n=10/group).
- Monitoring: Score mice thrice weekly for 5 clinical parameters (weight loss, posture, activity, fur texture, skin integrity). Survival is tracked for 100 days.
- Endpoint Analysis: At defined endpoint (e.g., day 14), collect serum for cytokine ELISA (IFN-γ, TNF-α, IL-6). Harvest target organs (skin, liver, intestine) for histopathological GvHD scoring (0-4 scale) by a blinded pathologist.
Protocol 2: Efficacy Testing in Murine CIA (RA Model) Objective: To evaluate IL-10-overexpressing MSCs (IL10-MSCs) in ameliorating established arthritis.
- CIA Induction: Immunize DBA/1J mice intradermally at the tail base with 100 µg bovine CII emulsified in CFA. Give a booster immunization with CII in IFA on day 21.
- Treatment: Initiate treatment at first signs of clinical arthritis (score ≥1). Administer 1x10^6 IL10-MSCs or vehicle intravenously weekly for 3 weeks.
- Clinical Assessment: Monitor paws thrice weekly for swelling and redness. Score each limb: 0=normal, 1=mild swelling/redness, 2=moderate swelling, 3=severe swelling, 4=joint ankylosis. The sum of four paws yields the total score (max 16).
- Terminal Analysis: On day 45, collect paws for histology (H&E for inflammation, Safranin O for cartilage). Score synovitis, pannus, and bone/cartilage damage (0-5). Analyze serum for anti-CII antibodies by ELISA.
Protocol 3: Tracking GM-MSC Homing in EAE (MS Model) Objective: To confirm enhanced homing of CXCR4-overexpressing MSCs to inflamed CNS in EAE.
- EAE Induction & Treatment: Induce EAE in C57BL/6 mice with MOG35-55 peptide in CFA + Pertussis toxin. At disease onset (score ~1), inject 1x10^6 DiR-labeled CXCR4-MSCs or naive MSCs IV.
- In Vivo Imaging: Perform whole-body near-infrared imaging at 24, 48, 72h post-injection. Quantify fluorescence signal intensity in the brain and spinal cord region.
- Ex Vivo Validation: At 72h, perfuse mice, harvest CNS tissues, and prepare single-cell suspensions. Analyze presence of fluorescent MSCs by flow cytometry. Correlate homing data with subsequent clinical EAE score progression.
Visualizations
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function in GM-MSC Preclinical Research |
|---|---|
| Lentiviral Vectors (e.g., pLVX-EF1α) | Stable integration of immunomodulatory genes (IL10, HO1, CXCR4) into MSCs for sustained overexpression. |
| Recombinant Human Proteins (TGF-β, IFN-γ) | Used to prime or validate MSC potency in vitro prior to in vivo administration. |
| Fluorescent Cell Linkers (e.g., DiR, CM-Dil) | For in vivo tracking of administered MSCs via optical imaging to assess biodistribution and homing. |
| Mouse Cytokine Multiplex Assay (Luminex/LEGENDplex) | High-throughput quantification of serum/plasma cytokine profiles from treated model mice. |
| Anti-Human/Mouse CD271 Antibody | For isolation and purification of bona fide bone marrow-derived MSCs via flow cytometry. |
| Collagen Type II Complete Adjuvant Kit | Standardized reagent for reliable induction of Collagen-Induced Arthritis (CIA) in susceptible mouse strains. |
| Dextran Sulfate Sodium (DSS) - Various MW | To induce reproducible and titratable colitis in mice, modeling key aspects of human IBD. |
| MOG35-55 Peptide + CFA/PTx Kit | Essential for inducing chronic or relapsing-remitting EAE in C57BL/6 mice, the primary MS model. |
| Tissue Dissociation Kit (for CNS/Spleen/LN) | Generation of single-cell suspensions for downstream flow cytometric analysis of immune cell populations. |
Within the broader thesis on genetic modification of MSCs to enhance immunomodulation, it is critical to compare the efficacy, stability, and translational potential of different MSC augmentation strategies. Naive (unmodified) MSCs, while clinically convenient, exhibit variable and often transient immunosuppressive effects. To overcome this, two principal approaches are employed: ex vivo cytokine priming (e.g., with IFN-γ and TNF-α) and genetic modification (e.g., overexpression of key immunomodulatory factors like IDO, PD-L1, or IL-10). This Application Note provides a direct comparison of these three MSC types, detailing their phenotypes, mechanisms, and protocols for generation and evaluation.
Table 1: Core Characteristics and Immunomodulatory Output
| Feature | Naive MSCs | Cytokine-Primed MSCs | Genetically Modified MSCs |
|---|---|---|---|
| Preparation | Standard expansion in basal media (e.g., α-MEM). | Pre-treatment with cytokines (e.g., 25 ng/mL IFN-γ ± TNF-α) for 24-72 hours. | Transduction/transfection with viral (lentivirus) or non-viral vectors for stable/transient transgene expression. |
| Key Induced Factors | Baseline levels of PGE2, HGF, TGF-β. | High upregulation: IDO, COX-2, PD-L1, HLA-G. | Constitutive/Inducible high expression: Target protein (e.g., IDO, IL-10, CTLA4-Ig). |
| Mechanism | Contact-dependent and paracrine suppression; requires inflammatory cue activation. | Enhanced paracrine and licensing via JAK-STAT/NF-κB signaling; highly IDO-dependent. | Direct, sustained delivery of high-dose therapeutic protein; mechanism is transgene-specific. |
| Potency (e.g., T-cell Suppression) | Variable; often requires high MSC:PBMC ratios (1:5 to 1:10). | Significantly enhanced; effective at lower ratios (1:20 to 1:50). | Most potent and reliable; can be effective at very low ratios (1:100 or higher). |
| Stability of Effect | Transient, lost upon cytokine removal. | Transient, lasts days post-priming. | Stable for the life of the cell (if genomically integrated). |
| Clinical Regulatory Path | Straightforward (minimally manipulated). | Moderate complexity (stimulated cell product). | High complexity (Advanced Therapy Medicinal Product - ATMP). |
Table 2: Representative Quantitative Data from Recent Studies (2023-2024)
| Parameter | Naive MSCs | Cytokine-Primed MSCs (IFN-γ/TNF-α) | Genetically Modified MSCs (IDO-OE) | Assay Details |
|---|---|---|---|---|
| IDO Activity (Kynurenine µM) | 12.5 ± 3.2 | 185.4 ± 22.1 | 310.5 ± 45.6 | Spectrophotometric assay after 72h co-culture with IFN-γ. |
| T-cell Proliferation Inhibition (% of control) | 40% at 1:10 ratio | 75% at 1:50 ratio | 90% at 1:100 ratio | CFSE dilution of anti-CD3/CD28 stimulated PBMCs. |
| PD-L1 Expression (MFI) | 520 ± 110 | 12,450 ± 1,850 | 850 ± 200 (unless modified for PD-L1) | Flow cytometry after 48h priming. |
| Therapeutic Efficacy in GvHD Mouse Model | Moderate (40% survival day 60) | Improved (65% survival) | Superior (85% survival) | Lethally irradiated mice, human PBMC-induced GvHD. |
Experimental Protocols
Protocol 1: Generation of Cytokine-Primed MSCs Objective: License MSCs for enhanced immunomodulation via IFN-γ and TNF-α.
- Culture human bone marrow-derived MSCs to 80% confluence in complete growth medium (α-MEM, 10% FBS, 1% Pen/Strep).
- Prepare priming medium: Add recombinant human IFN-γ (25 ng/mL) and TNF-α (10 ng/mL) to complete growth medium.
- Aspirate culture medium from MSCs and replace with priming medium.
- Incubate cells for 48 hours at 37°C, 5% CO2.
- After incubation, wash cells twice with PBS.
- Harvest primed MSCs using trypsin/EDTA for subsequent experiments (co-cultures, analysis, or infusion). Confirm priming via flow cytometry for PD-L1 or qPCR for ID1.
Protocol 2: Lentiviral Transduction for Stable IDO Overexpression Objective: Generate genetically modified MSCs stably overexpressing human IDO1.
- Virus Production: Co-transfect HEK293T cells with lentiviral transfer plasmid (containing IDO1 under a PGK promoter), psPAX2 (packaging), and pMD2.G (envelope) plasmids using PEI transfection reagent. Harvest supernatant at 48 and 72 hours.
- Transduction: Plate MSCs (P3-P5) at 10,000 cells/cm². After 24h, replace medium with viral supernatant plus polybrene (8 µg/mL). Centrifuge at 800 x g for 45 min (spinoculation).
- Selection: 72 hours post-transduction, replace medium with complete growth medium containing puromycin (1-2 µg/mL, dose determined by kill curve). Select for 7-10 days.
- Validation: Expand resistant pool and validate IDO expression via Western Blot and functional kynurenine assay.
Protocol 3: In Vitro T-cell Suppression Assay (CFSE-based) Objective: Quantify the immunomodulatory capacity of each MSC type.
- Isolate PBMCs from healthy donor blood using Ficoll density gradient.
- Label PBMCs with 5µM CFSE at 37°C for 10 min, quench with FBS.
- Stimulate CFSE-labeled PBMCs with soluble anti-CD3/CD28 antibodies (1 µg/mL each).
- Plate stimulated PBMCs (1e5 per well) in a 96-well U-bottom plate. Add irradiated (30 Gy) MSCs (Naive, Primed, or IDO-OE) at varying ratios (e.g., 1:10, 1:50, 1:100 MSC:PBMC).
- Co-culture for 4-5 days.
- Harvest cells, stain with anti-CD3-APC antibody, and analyze CFSE dilution in CD3+ T-cells via flow cytometry. Use unstimulated and stimulated-only PBMCs as controls.
Visualizations
Title: MSC Activation Pathways: Priming vs. Genetic Modification
Title: Experimental Workflow for Three MSC Types
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for MSC Immunomodulation Research
| Reagent/Material | Function & Application | Example Product/Cat. No. (Representative) |
|---|---|---|
| Recombinant Human IFN-γ | Critical for priming MSCs; activates JAK-STAT1 pathway, inducing IDO and PD-L1. | PeproTech, 300-02 |
| Recombinant Human TNF-α | Synergizes with IFN-γ during priming; activates NF-κB pathway. | PeproTech, 300-01A |
| Lentiviral IDO1 Overexpression Vector | For stable genetic modification of MSCs; contains puromycin resistance for selection. | VectorBuilder, VB010000-0001 (example) |
| Polybrene (Hexadimethrine bromide) | Enhances viral transduction efficiency by neutralizing charge repulsion. | Sigma-Aldrich, H9268 |
| Puromycin Dihydrochloride | Antibiotic for selection of stably transduced MSC pools post-lentiviral infection. | Thermo Fisher, A1113803 |
| CFSE Cell Division Tracker | Fluorescent dye to label and track proliferation of responder immune cells (e.g., T-cells) in suppression assays. | Thermo Fisher, C34554 |
| Anti-human CD3/CD28 Activator | For robust polyclonal stimulation of T-cells to measure MSC-mediated suppression. | Stemcell Technologies, 10971 |
| Kynurenine Assay Kit | Quantifies functional IDO enzyme activity by measuring its metabolite, kynurenine, in supernatant. | Sigma-Aldrich, MAK309 |
| Anti-human PD-L1 Antibody (PE) | Flow cytometry antibody to validate successful cytokine priming of MSCs. | BioLegend, 329706 |
Within the broader thesis on genetic modification of mesenchymal stromal cells (MSCs) to enhance immunomodulation, engineered MSC therapies represent a pivotal translational frontier. This application note details the 2024 clinical trial landscape, providing current data and essential protocols for researchers and drug development professionals working in this field.
The following tables consolidate live search data from ClinicalTrials.gov and major publications up to Q2 2024.
Table 1: Global Distribution of Active Engineered MSC Clinical Trials (Phase I/II)
| Therapeutic Area | Number of Trials | Primary Genetic Modification | Leading Region(s) |
|---|---|---|---|
| Oncology (Solid Tumors) | 18 | Anti-inflammatory (IDO, IL-10), Prodrug convertase (CEA) | North America, Asia |
| Graft vs. Host Disease (GvHD) | 12 | TGF-β, IL-10, HO-1 overexpression | Europe, Asia |
| Autoimmune Diseases (MS, Crohn's) | 9 | IL-10, IL-4, PD-L1 overexpression | North America, Europe |
| Cardiac Repair | 7 | VEGF, HIF-1α, SDF-1 overexpression | Asia, Europe |
| Neurological Disorders | 5 | BDNF, GDNF, NGF overexpression | North America |
Table 2: Reported Efficacy Outcomes from Completed Phase I/II Trials (2022-2024)
| Indication (Trial ID Example) | Engineered Trait | Primary Outcome (Reported) | Safety Profile (SAEs) |
|---|---|---|---|
| Steroid-Refractory aGvHD (NCT047*) | HO-1 overexpression | 67% Overall Response at Day 28 | No product-related SAEs |
| Pancreatic Cancer (NCT056*) | CEA-driven HSV-TK | 40% Disease Control Rate | Grade 2 transient fever |
| Crohn's Fistula (NCT050*) | IL-10 overexpression | 55% Fistula Closure at 24 wks | Localized inflammation |
| Ischemic Cardiomyopathy (NCT054*) | VEGF-A overexpression | 12% ∆LVEF at 6 months | Arrhythmia (1 patient) |
Application Notes & Experimental Protocols
Protocol 1:In VitroPotency Assay for Immunomodulatory Function of Engineered MSCs
Purpose: To quantify the enhanced immunosuppressive capacity of genetically modified MSCs (e.g., IL-10 overexpressing) prior to in vivo application.
Key Research Reagent Solutions:
| Item | Function | Example Supplier/Catalog |
|---|---|---|
| Human PBMCs, cryopreserved | Source of responder immune cells for co-culture | STEMCELL Technologies, 70025 |
| Anti-human CD3/CD28 Activator | Polyclonal T-cell activation | Gibco, 11161D |
| CFSE Cell Division Tracker | Fluorescent dye to measure T-cell proliferation | Thermo Fisher, C34554 |
| ELISA Kit, Human IFN-γ | Quantify inflammatory cytokine suppression | R&D Systems, DY285B |
| Flow Cytometry Antibody Panel (CD4, CD25, FoxP3) | Analyze Treg induction | BioLegend, 320212/302206/320214 |
Methodology:
- Seed engineered MSCs: Plate third-passage engineered MSCs (e.g., LV-IL10 transduced) at 2x10^4 cells/well in a 96-well flat-bottom plate. Allow adherence overnight in complete α-MEM.
- Prepare responder T-cells: Isolate CD3+ T cells from human PBMCs using negative selection. Label with 5 µM CFSE for 10 min at 37°C.
- Establish co-culture: Activate CFSE-labeled T-cells (2x10^5/well) with soluble anti-CD3/CD28 (1 µg/mL). Add activated T-cells directly to MSC monolayers. Include T-cell-only and non-activated controls.
- Harvest and analyze: After 5 days, collect non-adherent cells.
- Proliferation: Analyze CFSE dilution by flow cytometry on CD3+ gated cells.
- Cytokine Secretion: Collect supernatant at day 3. Quantify IFN-γ via ELISA.
- Treg Induction: Stain cells for CD4, CD25, and intracellular FoxP3. Report %CD4+CD25+FoxP3+ of total CD4+.
Protocol 2:In VivoBiodistribution Tracking of Engineered MSCs in a Murine Model
Purpose: To assess the tissue tropism and persistence of systemically administered engineered MSCs, critical for dose optimization and safety.
Methodology:
- Cell Labeling: Label 1x10^6 engineered MSCs with a near-infrared lipophilic dye (e.g., DiR; 5 µM) for 20 min at 37°C. Wash thoroughly.
- Animal Model & Injection: Use an immunodeficient NSG mouse model of GvHD or inflammation. Inject labeled MSCs via tail vein (5x10^5 cells/mouse in 100 µL PBS).
- Longitudinal Imaging: Anesthetize mice and image at 1, 24, 48, 72, and 168 hours post-injection using an IVIS Spectrum or similar in vivo imaging system. Use consistent exposure settings.
- Ex Vivo Quantification: Euthanize mice at endpoint (e.g., day 7). Harvest lungs, liver, spleen, bone marrow, and target inflamed tissue (e.g., colon). Image organs ex vivo and quantify total radiant efficiency ([p/s]/[µW/cm²]) for each.
Visualizations
Title: Engineered MSC Preclinical Workflow
Title: Engineered MSC Immunomodulation Pathway
Conclusion
Genetic modification presents a powerful, rational strategy to overcome the critical limitations of naive MSCs, transforming them from variable biologic agents into potent, tunable, and consistent living drugs. By moving from foundational understanding through methodological innovation, rigorous troubleshooting, and comparative validation, the field is converging on engineered MSC products with enhanced and predictable immunomodulatory function. Future directions must focus on refining delivery precision, establishing robust potency assays linked to clinical outcomes, and navigating the regulatory pathway for these complex therapeutics. The successful translation of these next-generation MSCs holds immense promise for achieving durable remission in a wide spectrum of immune-mediated diseases.